Mimosa-AI
Health Pass
- License — License: Apache-2.0
- Description — Repository has a description
- Active repo — Last push 0 days ago
- Community trust — 16 GitHub stars
Code Fail
- eval() — Dynamic code execution via eval() in datasets/ScienceAgentBench/eval_programs/BBBC002_cell_count_eval.py
- eval() — Dynamic code execution via eval() in datasets/ScienceAgentBench/eval_programs/BurnScar_eval.py
- eval() — Dynamic code execution via eval() in datasets/ScienceAgentBench/eval_programs/CogSci_before_JNMF_extract_eval.py
- eval() — Dynamic code execution via eval() in datasets/ScienceAgentBench/eval_programs/CogSci_pattern_high_com_low_plot_eval.py
- eval() — Dynamic code execution via eval() in datasets/ScienceAgentBench/eval_programs/CogSci_pattern_high_sim_eval.py
- eval() — Dynamic code execution via eval() in datasets/ScienceAgentBench/eval_programs/CogSci_pattern_high_sim_plot_eval.py
- eval() — Dynamic code execution via eval() in datasets/ScienceAgentBench/eval_programs/EOF_standard_hgt_plot_eval.py
- eval() — Dynamic code execution via eval() in datasets/ScienceAgentBench/eval_programs/EOF_xarray_hgt_plot_eval.py
Permissions Pass
- Permissions — No dangerous permissions requested
This open-source framework is designed for autonomous scientific research. It automatically synthesizes and evolves multi-agent workflows to execute complex computational tasks, such as bioinformatics and data analysis, using MCP-based tool discovery.
Security Assessment
Overall Risk: Medium. The framework acts as a multi-agent system that generates and executes code to perform scientific tasks, which inherently involves system modifications and potential shell executions. While no dangerous permissions or hardcoded secrets were found, the codebase contains multiple instances of dynamic code execution via `eval()` located specifically within evaluation programs (`datasets/ScienceAgentBench/eval_programs/`). Although these appear to be limited to dataset evaluation scripts rather than the core operational framework, dynamic execution always increases risk. Users should be cautious of the prompts and datasets fed into the system to prevent potential arbitrary code execution.
Quality Assessment
The project is in active development (last push was today) and comes with the permissive Apache-2.0 license. It is backed by an academic institution (CNRS) and has an associated peer-reviewed paper, which provides a strong baseline of credibility. However, community trust and adoption are currently in the very early stages, evidenced by a low GitHub star count of 16.
Verdict
Use with caution: While well-documented and academically grounded, the tool is relatively new, has a small user base, and utilizes dynamic code execution methods that require strict environment controls.
Open-source Evolving AI-Framework for Autonomous Scientific Research (ASR). Evolve multi-agent workflows to perform computational scientific tasks.

Mimosa-AI 🌼🔬
An Open-source Evolving AI-Framework for Autonomous Scientific Research
Mimosa-AI 🌼 — like the mimosa plant that senses, learns, and adapts — Mimosa is an open-source framework for autonomous scientific research that automatically synthesizes task-specific multi-agent workflows and refines them through execution feedback. Built around MCP-based tool discovery, code-generating agents, and LLM-based evaluation, it offers academics a modular and auditable alternative to closed black-box systems.
Objectives:
- Reproduce scientific studies with traceability and rigor
- Automate computational scientific pipelines across domains such as bioinformatics, docking, metabolomics, and more
Citation: Mimosa Framework: Toward Evolving Multi-Agent Systems for Scientific Research
M. Legrand, T. Jiang, M. Feraud, B. Navet, Y. Taghzouti, F. Gandon, E. Dumont, L.-F. Nothias — arXiv:2603.28986, 2026 — DOI — [BibTeX]
Table of Contents
- How does it work?
- Example: Reproducing a Bioactive Molecular Networking Paper
- Prerequisites
- Installation
- Configuration
- Running Mimosa
- Workspace and Audit Trail
- Learning through Evolution of Multi-Agent Workflows
- Transparency
- Command Line Arguments
- Evaluation
- Phone Notifications
- Telemetry Setup
- License
- Citation
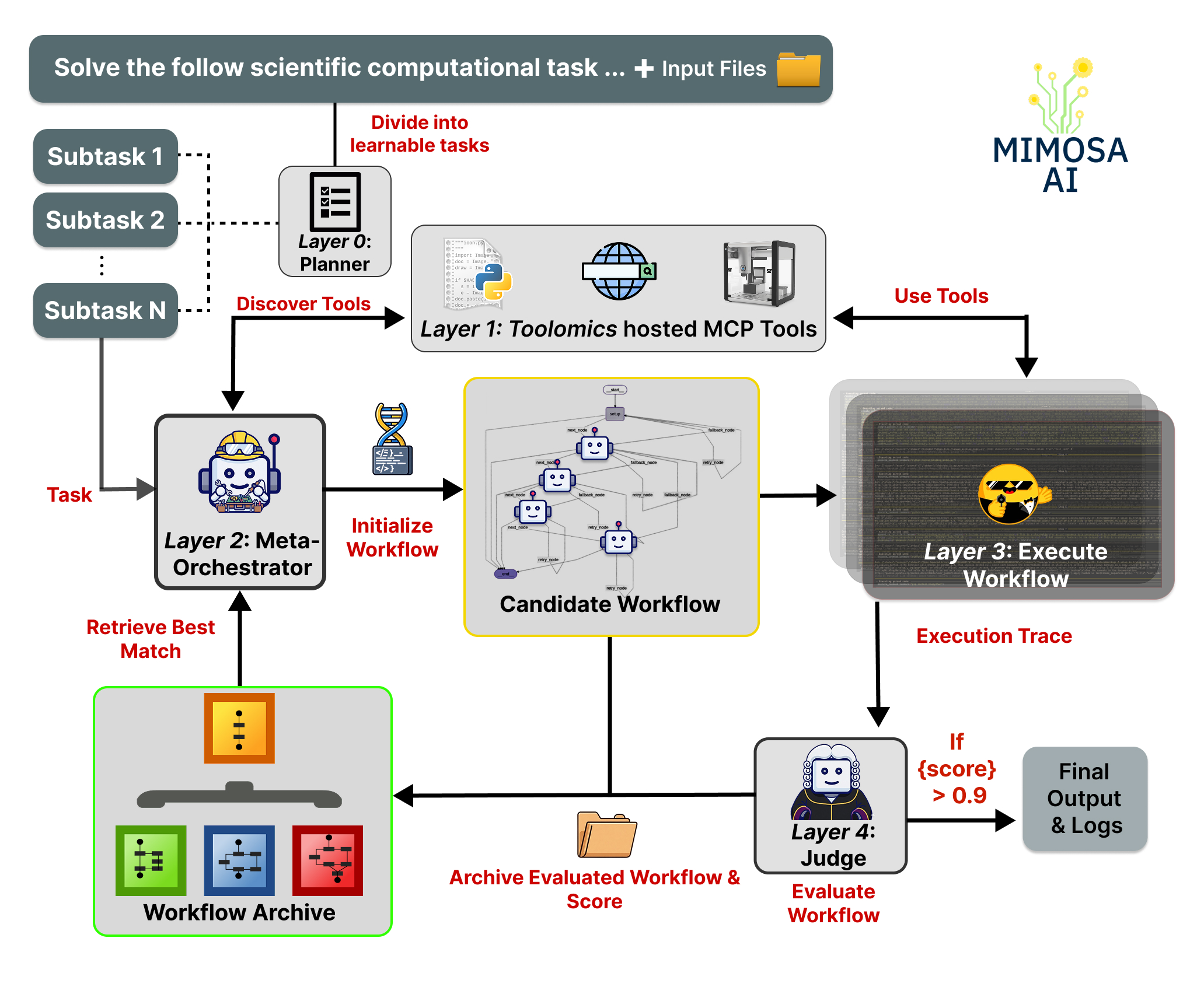
How does it work ?
The user gives Mimosa-AI either a high-level scientific goal or a single task.
- In goal mode, a planner decomposes the objective into smaller tasks. In task mode, the prompt is sent directly to the workflow synthesis layer.
- Mimosa discovers available MCP-based tools on the local network through Toolomics, from data-analysis utilities and web services to laboratory instruments.
- A meta-orchestrator synthesizes a task-specific multi-agent workflow and assigns relevant tools to specialized agents.
- Code-generating agents execute subtasks using discovered tools and scientific software libraries.
- An LLM-based judge evaluates execution traces and outputs; in learning mode, this feedback can drive iterative workflow refinement.
- Mimosa archives workflows, traces, logs, reports, and generated artifacts in a final capsule for inspection and reuse.
Example: Reproducing a Bioactive Molecular Networking Paper
Objective — Reproduce Nothias et al. (2018) end-to-end: from feature detection through network visualization, starting from
.mzMLfiles (raw data conversion assumed complete).
Mimosa-AI was given the paper, the raw data, and a set of domain-specific skills covering configuration details not fully specified in the publication.
Execution Timelapse
https://github.com/user-attachments/assets/dcd04ade-9c43-44a8-b3e3-a999d3dc895d
Output
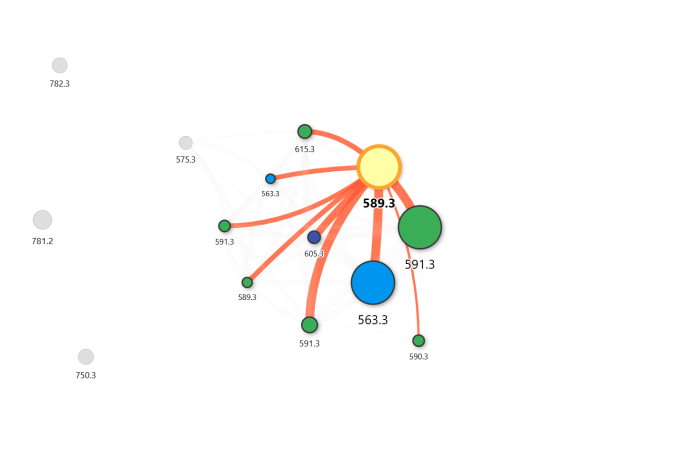
The resulting molecular network matches the topology reported in the original paper, including cluster separation and edge weights reproduced autonomously.
Prerequisites
- Python 3.10+
- uv (recommended) or pip
- A running Toolomics MCP server
Installation
1. Clone and create virtual environment
# Using uv (recommended)
pip install uv
uv venv .venv
source .venv/bin/activate # Windows: .venv\Scripts\activate
# Or with pip
python3 -m venv .venv
source .venv/bin/activate
2. Install dependencies
cd mimosa
uv pip install -r requirements.txt
3. Set API keys
Create a .env file at the project root. Include only the keys for the LLM providers you plan to use:
ANTHROPIC_API_KEY=... # Claude — recommended for workflow orchestration
OPENAI_API_KEY=... # OpenAI models - Optional
MISTRAL_API_KEY=... # Mistral models - Optional
DEEPSEEK_API_KEY=... # Deepseek - Optional
HF_TOKEN=... # HuggingFace provider, Optional
OPENROUTER_API_KEY=... # Any model via OpenRouter
# Optional — observability via Langfuse
LANGFUSE_PUBLIC_KEY=...
LANGFUSE_PRIVATE_KEY=...
4. Start the MCP server
Follow the setup instructions at HolobiomicsLab/toolomics. Configure it to run on a port range (e.g., 5000–5100).
Toolomics is Mimosa's companion platform for MCP server management. It exposes tools as discoverable MCP services, lets you register custom tools without modifying Mimosa's core orchestration logic, and provides the workspace where Mimosa reads and writes task artifacts. Both Mimosa and Toolomics are released under the Apache License 2.0.
Custom MCP tools can be added via the Toolomics docs.
Configuration
cp config_default.json my_config.json
Edit my_config.json. Key parameters:
| Parameter | Description |
|---|---|
workspace_dir |
Path to the Toolomics workspace — all generated files appear here |
discovery_addresses |
IP + port ranges for MCP server discovery |
planner_llm_model |
LLM for task decomposition and planning |
prompts_llm_model |
LLM for workflow prompt generation |
workflow_llm_model |
LLM for multi-agent orchestration (recommended: anthropic/claude-opus-4-5 or z-ai/glm-5) |
smolagent_model_id |
Model for SmolAgents execution subtasks |
judge_model |
LLM for output self-evaluation and scoring |
learned_score_threshold |
Minimum score to accept a result and stop iterating |
max_learning_evolve_iterations |
Maximum self-improvement iterations before accepting the result |
Running Mimosa
Mimosa supports two execution modes: Goal and Task.
Goal mode — multi-step scientific objective
Use this when your objective requires planning across multiple distinct operations (e.g., reproducing a paper, building an ML pipeline).
uv run main.py --goal "Your scientific objective" --config my_config.json
Examples:
uv run main.py \
--goal "Reproduce experiments from 'Dual Aggregation Transformer for Image Super-Resolution' (https://arxiv.org/pdf/2306.00306) and compare results." \
--config my_config.json
uv run main.py \
--goal "Develop a machine learning model to predict protein-ligand binding affinity." \
--config my_config.json
Task mode — single granular operation
Use this for a focused, self-contained operation without long-term planning.
uv run main.py --task "Your task description" --config my_config.json
Examples:
uv run main.py \
--task "Train a multitask model on the Clintox dataset to predict drug toxicity and FDA approval status." \
--config my_config.json
uv run main.py --task "Conduct a literature review on graph neural networks for drug discovery." --config my_config.json
Benchmark note: The results reported in the manuscript are measured in
taskmode, with the planning layer disabled, to isolate workflow synthesis and iterative refinement.Note: Toolomics must be installed and the MCP server must be running before executing any mode.
Workspace and Audit Trail
During execution, Mimosa reads and writes files inside the Toolomics workspace configured by workspace_dir. When a run finishes, the workspace contents are copied into a timestamped folder under runs_capsule/ so the final state is preserved as an archive rather than remaining only in a transient working directory.
- Toolomics
workspace/: the live working directory for the current run, including intermediate files, scripts, downloads, and generated outputs. sources/workflows/<uuid>/: the generated workflow and its execution metadata, includingstate_result.json,evaluation.txt,reward_progress.png, andmemory/traces.runs_capsule/<capsule_name>/: the archived snapshot of the run, copied from the Toolomics workspace for later inspection, comparison, or sharing.memory_explorer.py <uuid>: replay a workflow execution step by step to inspect agent traces, tool calls, and outputs.
Together, these locations form Mimosa's audit trail: they preserve what the system planned, executed, evaluated, and produced, supporting debugging, inspection, and potential replication.
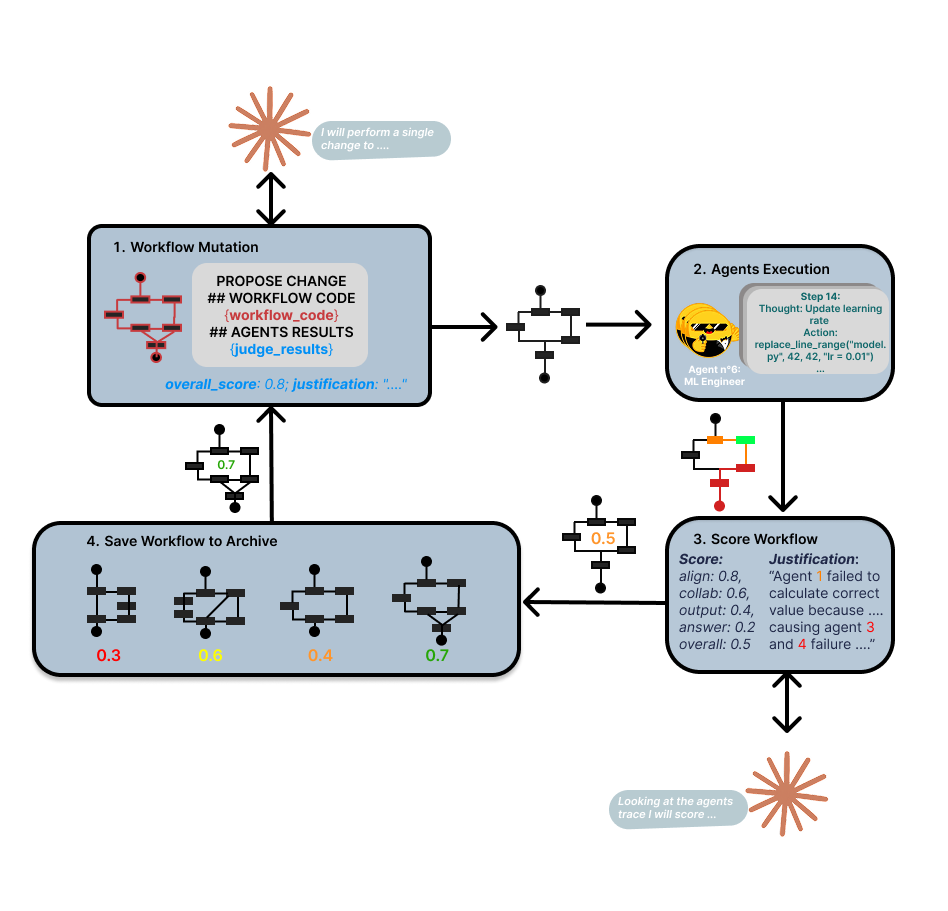
Learning through Evolution of Multi-Agent Workflows
Mimosa-AI is a self-evolving multi-agent system that dynamically synthesizes specialized workflows for scientific tasks. Rather than forcing tasks through fixed pipelines, the system composes custom multi-agent architectures on-demand and learns from execution patterns to optimize future performance.
The full architecture is organized into five layers: (0) optional planning, (1) MCP-based tool discovery, (2) meta-orchestration, (3) agent execution, and (4) judge/evaluation. In benchmark task mode, the planning layer is bypassed so workflow synthesis and refinement can be evaluated directly.
Mimosa is also capable of learning from failures through Darwinian-inspired evolution of multi-agent workflows. For each task, it composes a custom multi-agent graph and refines it through single-incumbent local search: at each iteration, only the best-performing workflow generates a successor, and only improvements are kept. Over time, the system builds a library of proven workflows, so similar future tasks start from a strong baseline rather than from scratch.
For any new task, start with learn mode to let it build competence before full autonomy.
Start in Learning mode
uv run main.py --task "Train a multitask model on the Clintox dataset to predict drug toxicity and FDA approval status" --learn --config my_config.json
Progress visualization:
Once Mimosa-AI completes its learning phase on a task, you can visualize exactly how it improved over time. The reward progress plot showing performance gains across attempts is automatically saved.
The reward progress plot is saved under the sources/workflows/<uuid> folder under the filename reward_progress.png.
Example:
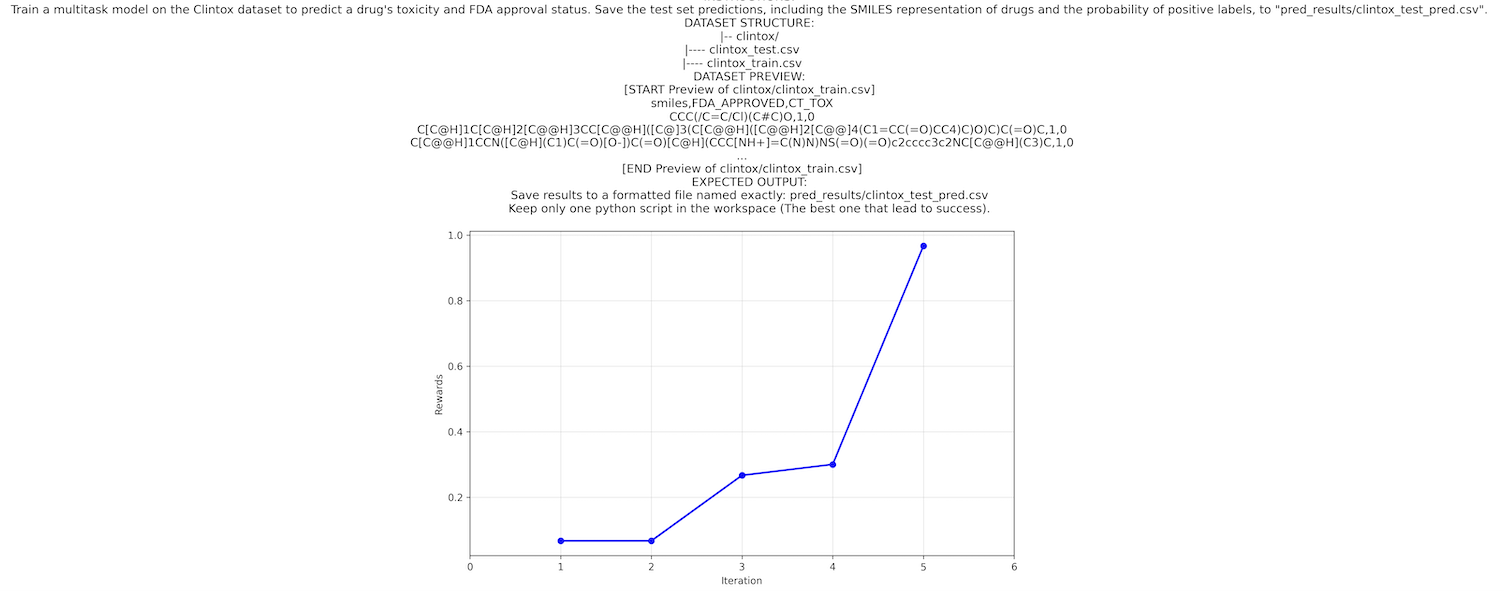
Transparency
We ship an interactive debugger, memory_explorer.py, that lets you step through any agent execution in granular detail.
Start it with a workflow (eg: memory_explorer.py 20260115_113303_9bb63437)
This replays the full execution trace—thoughts, tool calls and outputs so you can inspect exactly how decisions unfolded.
Command Line Arguments
Execution Modes
| Argument | Description |
|---|---|
--goal GOAL |
Specify a high-level research objective, paper reproduction, or scientific question (planner mode) |
--task TASK |
Execute a single task: literature review, datasets download, implement a machine learning model... |
--manual |
Interactive CLI mode to debug MCPs and test Mimosa tools directly |
--papers <CSV path> |
Evaluation on a CSV dataset containing research papers and prompts |
--science_agent_bench |
Evaluation on ScienceAgentBench |
Other parameters
| Argument | Description |
|---|---|
--learn |
Enable iterative-learning to optimize task performance |
--max_evolve_iterations N |
Maximum learning iterations |
--csv_runs_limit N |
Limit number of CSV entries to evaluate |
--scenario <scenario file name> |
Use specific scenario based assertions instead of LLM-as-a-judge for scoring execution |
--single_agent |
Single agent mode. fast, but can't improve throught learning |
--debug |
Enable debug mode for more verbose logging |
Evaluation
Mimosa-AI can be evaluated either on ScienceAgentBench or PaperBench.
⚠️ For unbiased evaluation it is advised to run ./cleanup.sh first, this will prevent Mimosa from using existing or cached workflows.
ScienceAgentBench
Manuscript-aligned results snapshot (task mode, 102 tasks):
- DeepSeek-V3.2 single-agent:
SR 38.2%,CBS 0.898,$0.05/task - DeepSeek-V3.2 one-shot multi-agent:
SR 32.4%,CBS 0.794,$0.38/task - DeepSeek-V3.2 iterative-learning:
SR 43.1%,CBS 0.921,$1.7/task - We also report model-dependent behavior in the manuscript: iterative learning improves GPT-4o, but yields marginal degradation for Claude Haiku 4.5.
To evaluate on ScienceAgentBench you must:
- download the ScienceAgentBench full dataset:
dataset link - unzip it with password:
scienceagentbench - copy content of
benchmark/benchmark/datasetsfolder toMimosa-AI/datasets/scienceagentbench/datasets
Evaluation on ScienceAgentBench with learning
uv run main.py --science_agent_bench --learn
Evaluation on ScienceAgentBench limited to 10 tasks with learning limited to 4 learning iterations
uv run main.py --science_agent_bench --csv_runs_limit 10 --max_evolve_iterations 4
PaperBench
Evaluation on OpenAI PaperBench with learning mode
OpenAI PaperBench is a benchmark for evaluating the ability of AI agents to replicate AI research, from the paper PaperBench: Evaluating AI’s Ability to Replicate AI Research.
uv run main.py --papers datasets/paper_bench.csv --csv_runs_limit 20 --learn
⚠️ This will save in runs_capsule/ folder the result of all paper's reproduction attempt, refer to Paper Bench documentation for complete evaluation.
Evaluation on custom benchmark of research paper
Place your benchmark CSV with the same format as
paper_bench.csvindatasets/<your_benchmark_name>.csv.Run on your benchmark:
uv run main.py --papers datasets/<your_benchmark_name>.csv --csv_runs_limit 20 --learn
Phone Notifications
Receive real-time updates about Mimosa's status via Pushover notifications.
Setup Instructions
Create Pushover Account
- Visit pushover.net
- Register and note your User Key
Create Application
- In Pushover dashboard, click "Create an Application/API Token"
- Name it "Mimosa" and copy the generated API Token
Configure Environment
export PUSHOVER_USER="your_user_key" export PUSHOVER_TOKEN="your_api_token"Install Mobile App
- Download Pushover from your device's app store
- Log in with your Pushover account
Telemetry Setup
Monitor and debug AI agents with real-time observability dashboards using Langfuse.
Quick Start
Deploy Langfuse Locally
git clone https://github.com/langfuse/langfuse.git cd langfuse docker compose up -dConfigure Environment Variables
Add to your
.envfile:LANGFUSE_PUBLIC_KEY=your_public_key LANGFUSE_PRIVATE_KEY=your_private_keyAccess Dashboard
While Mimosa-AI is running, visit
http://localhost:3000
Available Metrics
The telemetry dashboard provides:
- Agent Execution Traces: Step-by-step workflow visualization
- Performance Metrics: Response times and success rates
- Error Debugging: Detailed failure analysis
- Resource Usage: Token consumption and API calls
Example Dashboard: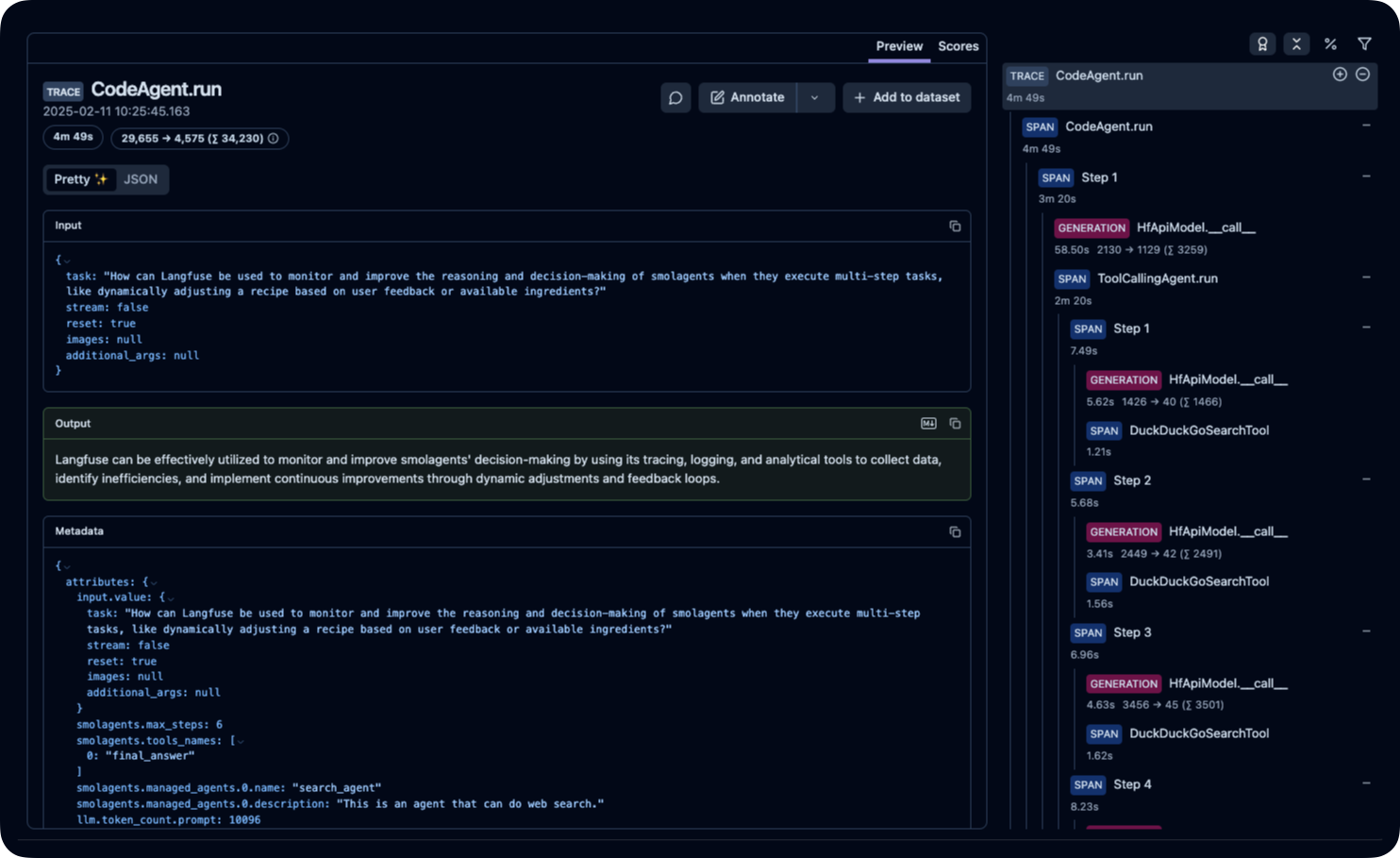
Note: Telemetry is optional but recommended for debugging and performance optimization.
License
This repository is publicly distributed under the Apache License 2.0. Apache 2.0 is a permissive open-source license that allows commercial use, modification, and redistribution, provided that redistributions preserve the applicable license, copyright, patent, trademark, attribution, and NOTICE notices and that modified files carry prominent notices stating that changes were made.
For contribution and licensing details, see:
NOTICEdocs/licensing-notes.mdCLA/INDIVIDUAL_CLA.mdCLA/EMPLOYER_AUTHORIZATION.md
Citation
If you find this work useful, please cite:
@article{legrand2026mimosa,
title={Mimosa Framework: Toward Evolving Multi-Agent Systems for Scientific Research},
author={Legrand, Martin and Jiang, Tao and Feraud, Matthieu and Navet, Benjamin and Taghzouti, Yousouf and Gandon, Fabien and Dumont, Elise and Nothias, Louis-F{\'e}lix},
journal={arXiv preprint arXiv:2603.28986},
year={2026}
}
Reviews (0)
Sign in to leave a review.
Leave a reviewNo results found



