FORTHought
Health Warn
- No license — Repository has no license file
- Description — Repository has a description
- Active repo — Last push 0 days ago
- Low visibility — Only 7 GitHub stars
Code Pass
- Code scan — Scanned 12 files during light audit, no dangerous patterns found
Permissions Pass
- Permissions — No dangerous permissions requested
No AI report is available for this listing yet.
A locally-hosted AI research platform for Physics/STEM labs — MCP tool servers for spectroscopy, XRD, SEM, literature search, and OriginLab automation, built on Open WebUI with AMD ROCm.

|
FORTHoughtA locally hosted AI platform for physics and STEM laboratory workflows.



|
Paper: M. Adamidis, D. Katrisioti, Y. Tzitzikas, E. Stratakis, "It's not the Language Model, it's the Tool: Deterministic Mediation for Scientific Workflows" — arXiv:XXXX.XXXXX (2026). The paper evaluates how typed tool mediation produces identical scientific results across runs while code-generating approaches vary. The OriginMCP and SEM Micro servers in this repository are the tools evaluated in the paper.
What This Is
This repository contains the custom tools, MCP servers, functions, and configuration I built for a locally hosted AI research platform at a physics lab. It runs on Open WebUI and serves 15 active researchers across spectroscopy, electron microscopy, X-ray diffraction, photoluminescence, and general scientific workflows.
This is not a product. It is a working research platform that I maintain and iterate on. The tools sit on top of existing open-source projects; my contribution is in how they are assembled, configured, and extended for real lab use. I upload the code here for documentation, reproducibility, and to share with the community.
Hardware:
| Server | CPU | GPU | Role |
|---|---|---|---|
| Compute Server | AMD Ryzen 9 9950X | 2× AMD Radeon AI Pro R9700 (32 GB VRAM each) | OriginLab, Lemonade reranker, embeddings, VLM, LM Studio, Docling |
| Docker Server | Intel Xeon E5-2680 v4 | AMD Radeon RX 7900 XT (20 GB VRAM) | Open WebUI, MCP servers, Jupyter, Qdrant, MetaMCP, all Docker services |
The two machines communicate over Tailscale. The Docker server runs the full containerized stack (35+ containers), while the compute server handles GPU-intensive inference tasks.
Configuration note: All service URLs default to
localhost. To run your own instance, copyconfig/.env.exampleto.envand fill in your values.
Base Stack
- LLM inference via LM Studio (local models) and cloud APIs (Gemini via Google research grant, OpenRouter as fallback)
- Code execution via a GPU-accelerated Jupyter kernel (ROCm, scientific Python stack)
- Document parsing via Docling (native PyTorch ROCm, with Qwen3-VL for figure descriptions)
- Vector search via Qdrant (hybrid BM25 + semantic)
- Tool orchestration via MetaMCP, aggregating all MCP servers behind a single HTTP endpoint
All services run on-premises. No data leaves the local network unless a cloud LLM is explicitly used.
What I've Built
RAG Pipeline
Tuned Open WebUI's RAG pipeline for scientific documents. Out-of-the-box defaults struggle with multi-column papers, equations, and dense tables.
| Stage | Configuration | Notes |
|---|---|---|
| Parsing | Docling (ROCm GPU) | Granite-Docling + Qwen3-VL for figure descriptions |
| Embeddings | Qwen 0.6B embed via LM Studio | Served through a custom parallel proxy that splits batch requests into concurrent sub-batches |
| Vector store | Qdrant, hybrid search | 800-token chunks, 100 overlap |
| Reranking | BGE-reranker-v2-m3 (GGUF) on 🍋 AMD Lemonade | Shared by RAG and web search |
Multi-Role Local Models on Lemonade
Each locally hosted model serves multiple roles across the stack:
| Model | Roles | Used by |
|---|---|---|
| BGE-reranker-v2-m3 (GGUF) | RAG reranking, web search reranking | OWUI Documents pipeline, web_search.py tool |
| Qwen3-Embedding-0.6B (GGUF) | Document embeddings | OWUI RAG pipeline (via parallel proxy) |
| Qwen3-VL-30B (GGUF) | Image descriptions, SEM analysis, general VLM | Docling, image_description_context filter, SEM server |
Free Web Search with Reranking
Custom tool that provides web-augmented answers without expensive search APIs:
- Queries the LangSearch API (free tier, 1000 calls/day)
- Passes results through the same Lemonade reranker used by RAG
- Filters by relevance score, Latin-script ratio, and content length
- Auto-fetches top-scoring pages for full context
- Injects citations into OWUI's native citation UI
See: tools/web_search.py
Skill-Gated Tool Routing
Instead of one system prompt stuffed with every tool's documentation, the platform uses per-profile tool surfaces:
| Profile | Use case | Tools it sees |
|---|---|---|
| Lab | Literature, reports, web research, chemistry | paper.*, file.*, web.*, chem.* |
| Coder | Data analysis, plotting, scripting | Jupyter (ROCm), file.*, code tools |
| Instrument | Spectroscopy, SEM, XRD, PL | spec.*, micro.*, xrd.*, pl.* |
Each profile consists of:
- A lean system prompt (
profiles/) — routing table only, no tool recipes - A skills file (
skills/) — Python OWUI Tool acting as MCP gateway - Skill documents (
skill-docs/) — loaded on-demand only when the model actually needs a tool
This keeps token usage low. The model never loads documentation it doesn't need.
Science MCP Servers
These are the MCP tool servers built for the lab's workflows. Each runs as a standalone Python HTTP server implementing the MCP JSON-RPC protocol.
OriginMCP — OriginLab Automation
Server: mcp-servers/origin/server.py (3300+ lines)
Automates OriginPro through its COM API, allowing the model to inspect, analyze, and fit spectroscopy data from OPJ project files. This is the photoluminescence tool evaluated in the paper.
- Inspect OPJ file structure and column metadata (user-defined parameters: strain, polarization, power)
- Fit peaks: single or two-peak decomposition across Lorentzian, Gaussian, Voigt lineshapes
- Batch fit across N columns with automatic summary grid, waterfall, and trend plots
- Detect degenerate fits (converging peaks) and flag them without crashing
- Infer experimental parameters from column naming conventions when metadata is missing
Runs on Windows (requires OriginPro). Accessible to the Docker stack via Tailscale.
SEM Micro — Electron Microscopy FFT Analysis
Server: mcp-servers/micro/server.py
FFT-based periodicity and particle size analysis for SEM images. This is the scanning electron microscopy tool evaluated in the paper.
- Reads magnification from the SEM info bar via VLM
- 2D FFT with radial averaging and peak detection
- Reports periodicity (nm), orientation, confidence, and SNR for macro/micro frequency bands
- Particle size distribution (count, mean diameter, std dev, p10–p90)
- 6-panel composite figure output
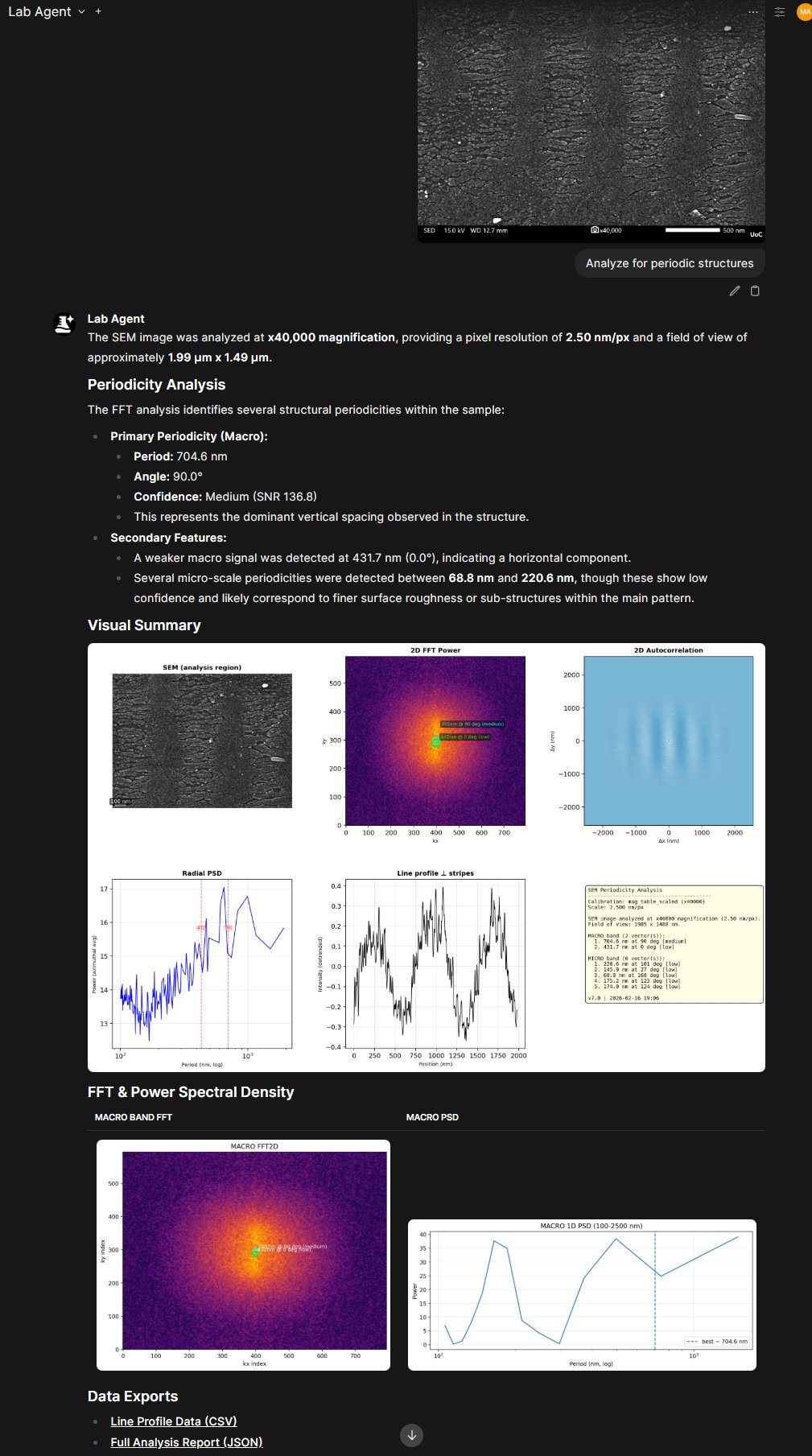
FFT periodicity analysis of a LIPSS nanostructure at ×40,000 — automatic magnification detection, 6-panel composite, and data export.
XRD Server — X-Ray Diffraction Phase Identification
Server: mcp-servers/xrd/server.py
File-format-agnostic XRD analysis pipeline that identifies crystalline phases from raw diffraction data.
- Parses
.xy,.dat,.csv,.brml(Bruker XML), and.raw(Bruker RAW v4 binary) - Matches against the Crystallography Open Database (COD) and Materials Project
- Reports confidence scores, Rwp R-factors, purity estimates, impurity detection
- Generates annotated pattern and comparison plots inline
- Exports Origin-ready CSVs
PL Server — Photoluminescence Experiment Planning
Server: mcp-servers/pl/server.py
Laser/filter/optics recommender and Fresnel interference calculator for PL measurements, with specialized support for 2D materials.
- Recommends laser wavelengths, filters, and optical components for any target material
- Calculates excitation and emission enhancement factors for air/SiO₂/Si stacks
- Models signal intensity variation with SiO₂ thickness (PL, Raman, SHG)
- Plans SHG/THG, strain analysis, PLE, imaging mode, and valley polarization experiments
- Built-in materials database (TMDs, perovskites, III-V semiconductors)
Papers MCP — Literature Search
Server: mcp-servers/papers/server.py
Searches and retrieves academic papers from multiple open-access sources.
- Search OpenAlex (250M+ papers, semantic search), PubMed, Semantic Scholar, NASA ADS, CrossRef, OpenAIRE
- Author disambiguation via ORCID
- PDF download from open-access sources (arXiv, OpenAlex, Unpaywall, PMC)
- Batch operations for reference lists
Files MCP — Document Generation
Server: mcp-servers/files/
Generates DOCX, PPTX, XLSX, PDF, CSV, HTML, and plain text files from structured data, with default templates.
📸 Screenshots: See the Examples Gallery for OriginMCP batch fitting, literature search, XRD phase ID, PL experiment planning, and more.
Custom Open WebUI Functions
Pipes
| Function | What it does |
|---|---|
gemini_pipe.py |
Google Gemini pipeline with native tool support and image generation |
openrouter_pipe_v2.py |
OpenRouter Responses API integration with web search and file uploads |
Note: The pipe files here are reference snapshots. The production pipes have evolved significantly with event-driven streaming, tool-call UI panels, and provider-specific optimizations.
Filters
| Function | What it does |
|---|---|
markdown_normalizer.py |
Cleans leaked reasoning labels, XML artifacts, and broken formatting from LLM outputs |
image_description_context.py |
Extracts text from images via a vision model so text-only models can process image uploads |
image_reembed_injector.py |
Re-injects uploaded image URLs into context for models that lose them across turns |
uploaded_filename.py |
Injects file metadata (IDs, names) so tools can reference uploaded files |
Tools
| Function | What it does |
|---|---|
web_search.py |
Free web search with LangSearch API + Lemonade reranking |
chemistry_database.py |
PubChem and CAS compound lookups (name, formula, safety data) |
chart_server/ |
Server-side Chart.js rendering for inline data visualizations |
presenton_adapter.py |
MCP bridge to Presenton for AI-generated slide decks |
Actions
| Function | What it does |
|---|---|
export_to_word.py |
Export conversations to formatted Word documents with APA 7th styling |
lemonade_control_panel.py |
Visual dashboard for monitoring the AMD Lemonade Server |
Architecture
┌─── Docker Server (Xeon E5-2680v4 + RX 7900 XT) ────────────┐
│ │
│ Open WebUI (:8081) │
│ │ │
│ ├── Gemini Pipeline (primary LLM) │
│ ├── OpenRouter Pipe (fallback models) │
│ │ │
│ ├── Qdrant (:6333) ── hybrid vector search │
│ ├── Jupyter (:8888) ── code execution (ROCm) │
│ │ │
│ └── MetaMCP (:12008) │
│ ├── Papers (:9005) ├── Chemistry (OWUI tool) │
│ ├── Files (:9004) ├── XRD (:9008) │
│ ├── SEM/Micro (:9006) ├── PL (:9010) │
│ └── Context7, Presenton, Browser │
│ │
└──────────────── Tailscale ───────────────────────────────────┘
│
┌─── Compute Server (Ryzen 9950X + 2× R9700 64 GB) ───────────┐
│ │
│ LM Studio (:1234) ── local LLMs, VLM (Qwen3-VL) │
│ Lemonade Server (:8040) ── BGE-reranker-v2-m3-GGUF │
│ Embedding Proxy (:5555) ── Qwen 0.6B embed │
│ Docling (:5001-5003) ── document parsing (ROCm) │
│ OriginMCP (:12009) ── OriginLab COM automation (Windows) │
│ │
└───────────────────────────────────────────────────────────────┘
Repository Structure
FORTHought/
├── README.md
├── CITATION.cff
├── LICENSE
├── .gitignore
│
├── mcp-servers/ # MCP tool servers
│ ├── papers/server.py # Literature search & download
│ ├── origin/server.py # OriginLab COM automation
│ ├── micro/server.py # SEM FFT periodicity analysis
│ ├── xrd/server.py # XRD phase identification
│ ├── pl/server.py # PL experiment planning
│ └── files/ # Document generation (DOCX, PPTX, etc.)
│
├── pipes/ # LLM provider integrations (reference snapshots)
│ ├── gemini_pipe.py # Google Gemini pipeline
│ └── openrouter_pipe_v2.py # OpenRouter Responses API
│
├── filters/ # Input/output processing
│ ├── markdown_normalizer.py
│ ├── image_description_context.py
│ ├── image_reembed_injector.py
│ └── uploaded_filename.py
│
├── tools/ # OWUI tool functions
│ ├── web_search.py
│ ├── chemistry_database.py
│ ├── chart_server/
│ └── presenton_adapter.py
│
├── actions/ # Chat action buttons
│ ├── export_to_word.py
│ └── lemonade_control_panel.py
│
├── skills/ # MCP gateway + routing logic
│ ├── lab_skills.py
│ ├── coder_skills.py
│ └── instrument_skills.py
│
├── skill-docs/ # On-demand tool documentation
│ ├── instruments/{origin,sem,xrd}/
│ ├── research/{academic-search,literature,web-search}/
│ ├── documents/{file-export,presenton}/
│ ├── compute/code-interpreter/
│ └── visualization/chartjs/
│
├── profiles/ # System prompt templates
│ ├── lab_prompt.md
│ ├── coder_prompt.md
│ └── instrument_prompt.md
│
├── config/
│ ├── docker-compose.yml
│ ├── .env.example
│ └── Dockerfile.*
│
├── docs/
│ ├── SETUP.md
│ └── EXAMPLES.md
│
├── scripts/
│ ├── fileserver_app.py
│ └── smoke_test_core.sh
│
└── assets/
├── forthought-logo.png
└── screenshots/
Getting Started
Clone:
git clone https://github.com/MariosAdamidis/FORTHought.git cd FORTHoughtConfigure:
cp config/.env.example .env # Edit .env with your API keys and service URLsStart:
docker compose -f config/docker-compose.yml up -dSet up Open WebUI: See docs/SETUP.md for importing functions, creating model profiles, configuring RAG, code execution, and MetaMCP wiring.
Verify:
bash scripts/smoke_test_core.sh
Roadmap
Done
- HTTP-persistent MCP transport
- Skill-gated multi-profile routing
- OriginMCP: batch fitting, waterfall/trend plots, degenerate fit detection
- XRD pipeline with COD/MP matching and Origin export
- PL substrate enhancement and experiment planning server
- Chemistry database integration
- AMD Lemonade reranker (dual: RAG + web search)
- Free web search tool with reranking
- Parallel embedding proxy
- Docling GPU acceleration on ROCm with Qwen3-VL
- Markdown normalizer for output cleanup
- Typed mediation pattern evaluated across 4 platforms (paper)
- Open WebUI 0.9.5 with custom patches
- Event-driven tool-call UI panels in production pipes
Next
- Additional OWUI tools to be open-sourced (interactive question UI, inline visuals, computer use)
- Browser-use VLM automation for instrument control
- N-peak fitting (defect emission, phonon replicas)
- Raman peak database
- PLE contour plots
- Full-loop: plan → measure → analyze → report
Security
This platform is designed for local/private-network deployment. Do not expose services directly to the public internet. See SECURITY.md for details.
The papers server in this repository is configured for open-access sources only. If you need institutional access, configure that through your own environment and credentials.
Acknowledgements
Built on top of open-source projects and community contributions.
Core Platform
- Open WebUI — Chat interface
- AMD Lemonade — Local reranker serving on AMD hardware
- Docling — Document parsing
- Qdrant — Vector database
- LM Studio — Local model serving
- MetaMCP — MCP server orchestration
- Crystallography Open Database — XRD reference patterns
- Materials Project — Materials science database
- LangSearch — Free web search API
Community Functions (adapted for FORTHought)
Several Open WebUI functions in this repository are adapted from community work. Original authors are credited in each file header.
| Function | Original Author | What I adapted |
|---|---|---|
| Gemini Pipeline | owndev, olivier-lacroix | Token optimization, tool integration, image generation |
| OpenRouter Pipe | rbb-dev | Integration with MetaMCP and file server |
| Export to Word | Fu-Jie | APA 7th Edition styling, Greek/English i18n |
| Markdown Normalizer | Fu-Jie | Custom rules for scientific output cleanup |
| Image Description Context | inMorphis | Adapted for multi-model routing |
| Files Metadata Injector | GlissemanTV | Integration with file server pipeline |
| Lemonade Control Panel | Sawan Srivastava | Integrated into Lemonade deployment |
Community Contributors
- r/LocalLLaMA contributor Ok_Ocelot2268 — ROCm patches for Unsloth
- @mballesterosc (Open WebUI community) — File Path Injector concept
Citation
If you use ideas or code from FORTHought, please cite:
@misc{adamidis2026forthought,
author = {Adamidis, Marios and Katrisioti, Danae and Tzitzikas, Yannis and Stratakis, Emmanuel},
title = {It's not the Language Model, it's the Tool: Deterministic Mediation for Scientific Workflows},
year = {2026},
eprint = {XXXX.XXXXX},
archivePrefix = {arXiv},
primaryClass = {cs.AI}
}
License
MIT License. See LICENSE for details.
Reviews (0)
Sign in to leave a review.
Leave a reviewNo results found