claude-paperloom
Health Pass
- License — License: Apache-2.0
- Description — Repository has a description
- Active repo — Last push 0 days ago
- Community trust — 43 GitHub stars
Code Pass
- Code scan — Scanned 12 files during light audit, no dangerous patterns found
Permissions Pass
- Permissions — No dangerous permissions requested
This tool is a Claude Code plugin that integrates with Obsidian to build a self-maintaining research knowledge graph. It allows users to input academic papers via URLs or PDFs, which are then automatically summarized, structured, and linked to related research in their local vault.
Security Assessment
Overall risk: Low. The automated code scan of 12 files found no dangerous patterns, no hardcoded secrets, and the tool does not request any dangerous system permissions. Because its primary function involves fetching and processing documents from sources like arXiv, it inherently makes outbound network requests. However, it operates locally within your file system and Obsidian vault. It does not appear to access unrelated sensitive data or execute hidden shell commands.
Quality Assessment
The project is in excellent health and actively maintained, with its most recent code push happening today. It is legally safe to use and build upon, covered by the standard and permissive Apache-2.0 license. Community trust is solid for a niche developer tool, currently backed by 43 GitHub stars. The documentation is clear, professional, and transparent about how the system processes academic data using LLM calls.
Verdict
Safe to use.
Claude Code Plugin for Self-maintaining research knowledge graph for Claude Code + Obsidian
A Claude Code Plugin for Self-maintaining research knowledge graph for Claude Code + Obsidian.
|

|
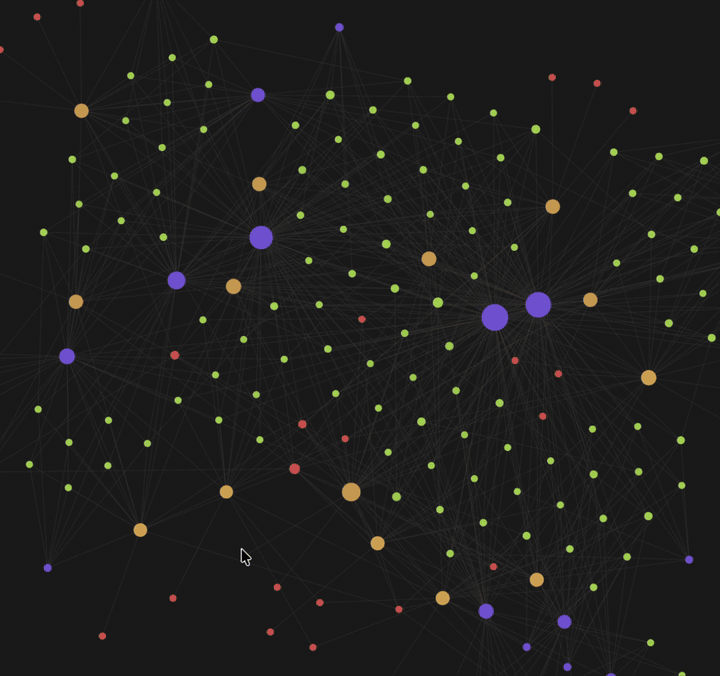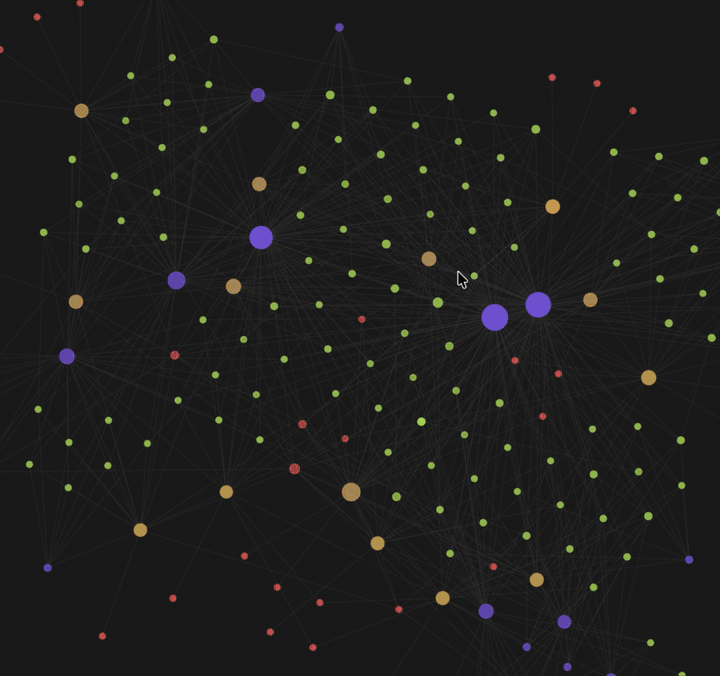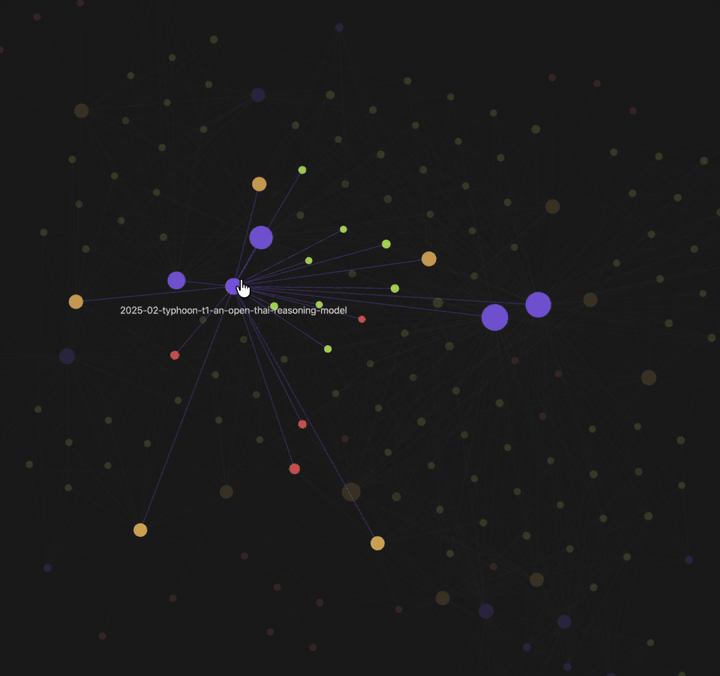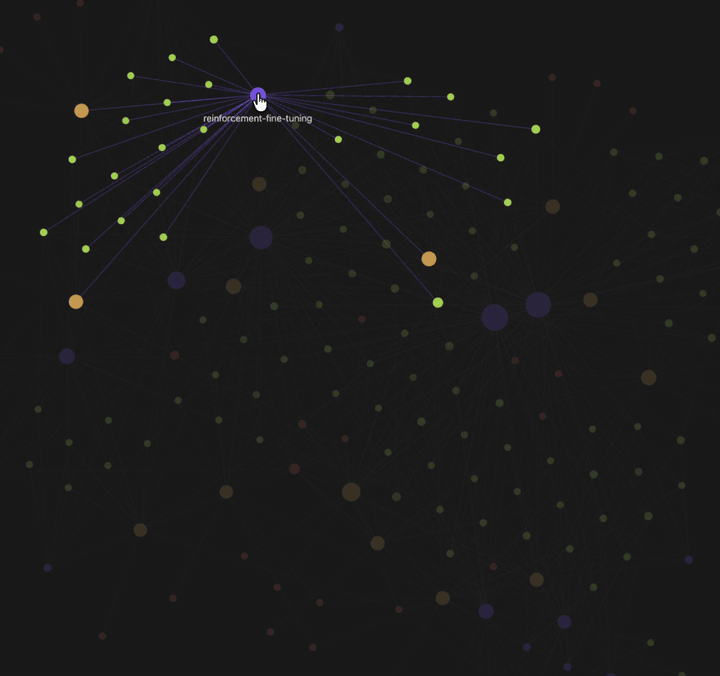
| Knowledge graph view — 🟡 papers · 🟢 findings · 🟣 fields · 🔴 authors. Example shown: Typhoon.AI research papers. | Per-paper info page — 4-section summary (incl. critique), atomic findings, metadata + quality scores, and typed edges to related work. |
Keep every paper you care about. See how they connect. Drop in a URL, arXiv ID, DOI, or PDF — Claude files it into an Obsidian vault, extracts the atomic claims, and wires them to everything you've read before with typed edges: supports, contradicts, extends, uses, similar-to.
Based on Andrej Karpathy's LLM Wiki pattern, tuned for research papers. 4 LLM calls per paper. Zero manual filing. Linking cost stays constant as the graph grows.
Features
- 4-section triage summary per paper — Key Takeaways → Background → Main Idea & Summary → Critique. The critique is generated by a reasoning model, so you get critical thinking, not a polite restatement of the abstract.
- Structured metadata + quality scores — authors, venue, date, fields, and a quality assessment (credibility, experimental rigor, reproducibility) on every paper.
- Atomic findings as graph nodes — one claim per file, with frontmatter and backlinks.
- Typed edges —
supports,contradicts,extends,uses,similar-to; auto-aggregated to paper-level (cites,builds-on, …). - Deterministic citation matching — arXiv ID → DOI → fuzzy title → first-author+year. The LLM never sees the reference list.
- Constant-cost linking — pre-filter caps the LLM at ≤30 candidate findings, so the 100th paper costs the same as the 10th.
- Cross-paper contradiction surfacing — bidirectional, with a pre-built Dataview view.
- Pre-built Dataview views — by-author, by-field, contradictions, high-credibility, recent-papers.
- Obsidian-native — plain wikilinks + YAML frontmatter, no custom app, no lock-in.
- Local model support — point at Ollama via
ANTHROPIC_BASE_URL.
Quick Start
# 1. Install the plugin
claude --plugin-dir /path/to/claude-paperloom
# 2. Set up Python deps (plugin expects a venv at the plugin root)
cd /path/to/claude-paperloom
python3.11 -m venv .venv
.venv/bin/pip install pyyaml pymupdf
# 3. Scaffold your vault (default ~/PaperLoom)
/paperloom:init
# 4. Ingest your first paper
/paperloom:ingest https://arxiv.org/abs/1706.03762
Open the vault folder in Obsidian and turn off Restricted Mode once to activate Dataview.
Commands
| You run | Claude does |
|---|---|
/paperloom:init [path] |
Scaffold the vault. Idempotent. Seeds Dataview. |
/paperloom:ingest <url|arxiv-id|doi|pdf> |
Fetch, summarize, extract findings, link. Skips if already in vault. |
/paperloom:query <question> |
Search + synthesize across the vault with wikilink citations. |
/paperloom:lint |
Health check. Auto-wires near-duplicate findings with bidirectional similar-to. |
Docs
- Pipeline — the 4-LLM-call ingest pipeline, section detection, models, token cost.
- Linking — five-layer linking, relation types, paper-level aggregation.
- Architecture — vault layout, plugin structure, design principles, out-of-scope.
Reviews (0)
Sign in to leave a review.
Leave a reviewNo results found


