ChatSpatial
Health Gecti
- License — License: MIT
- Description — Repository has a description
- Active repo — Last push 0 days ago
- Community trust — 26 GitHub stars
Code Gecti
- Code scan — Scanned 12 files during light audit, no dangerous patterns found
Permissions Gecti
- Permissions — No dangerous permissions requested
This tool is an MCP server that allows users to analyze spatial transcriptomics data through natural language. It replaces ad-hoc LLM code generation by exposing over 60 curated data analysis methods as accessible tools for AI clients like Claude or Codex.
Security Assessment
The overall risk is rated as Low. The automated code scan checked 12 files and found no dangerous patterns, hardcoded secrets, or requests for overly broad permissions. The tool processes local scientific data files (such as .h5ad formats) based on the file paths you provide, meaning you are in control of what data it accesses. Because it acts as a server orchestrating complex data science libraries, it inherently requires local execution and potentially outbound network requests to download underlying models or datasets, which is standard behavior.
Quality Assessment
The project demonstrates strong quality and maintenance signals. It is licensed under the permissive and standard MIT license. The repository is actively maintained (with the most recent push occurring today) and features continuous integration testing. It has solid community traction for a niche bioinformatics tool, currently sitting at 26 GitHub stars. Furthermore, the project appears to be backed by legitimate academic research, evidenced by published papers and presentations at major conferences like ICLR 2026.
Verdict
Safe to use.
🧬 Analyze spatial transcriptomics data through natural language conversation. Stop writing code, start having conversations with your data. MCP server for Claude Code and Codex.
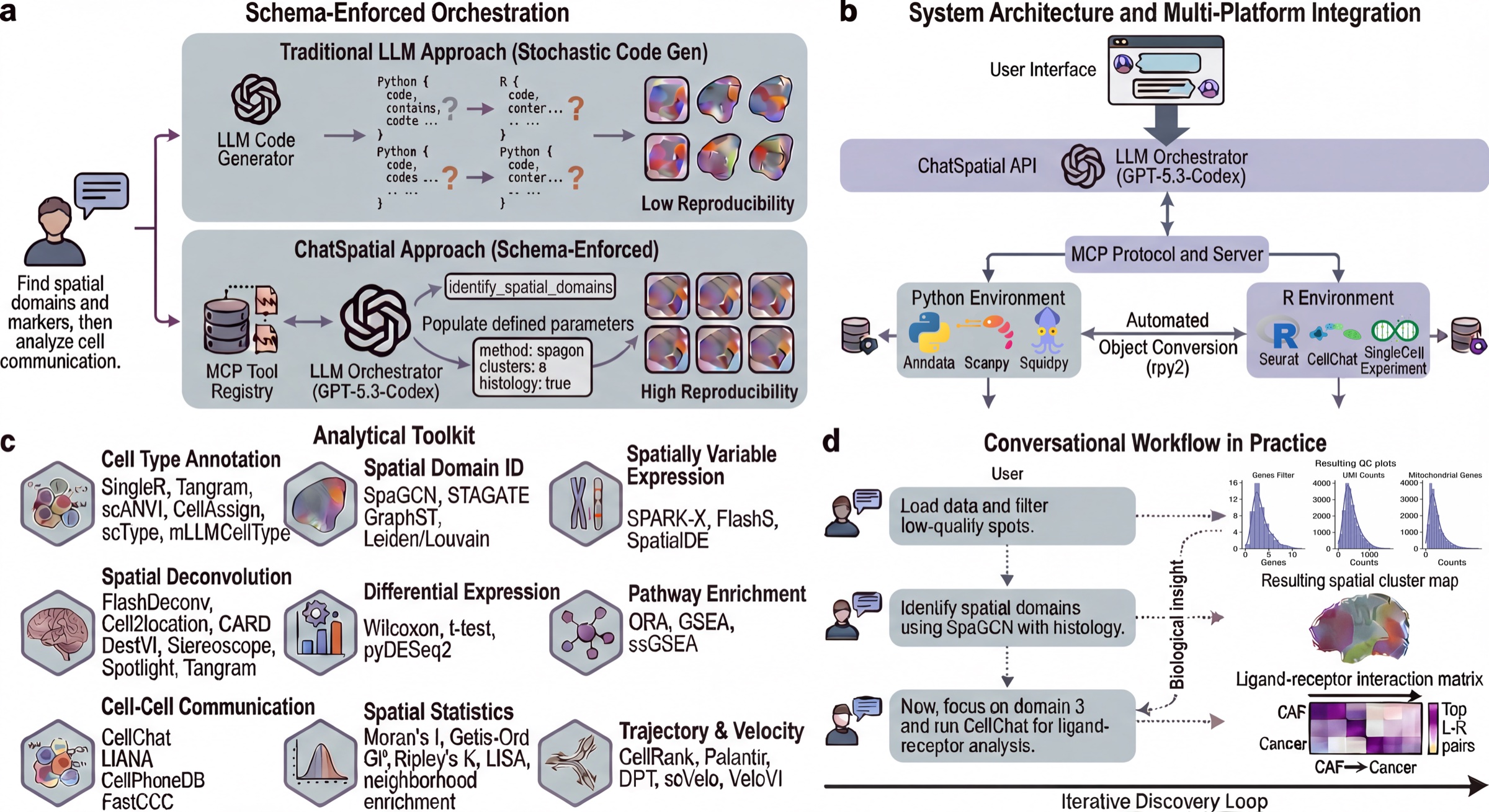
ChatSpatial replaces ad-hoc LLM code generation with schema-enforced orchestration. Instead of generating arbitrary scripts, the LLM selects tools and parameters from a curated registry, making spatial transcriptomics workflows more reproducible across sessions and clients.
It exposes 60+ spatial transcriptomics methods as MCP tools, so any MCP-compatible client can analyze data through natural language.
Start Here
- Install ChatSpatial — Installation Guide
- Configure your MCP client — Configuration Guide
- Run your first analysis — Quick Start
Minimal example prompt:
Load /absolute/path/to/spatial_data.h5ad and show me the tissue structure
ChatSpatial works with any MCP-compatible client — Claude Code, Claude Desktop, Codex, OpenCode, and other MCP-capable tools.
Capabilities
60+ methods across 11 categories. Supports 10x Visium, Xenium, Slide-seq v2, MERFISH, seqFISH.
| Category | Methods |
|---|---|
| Spatial Domains | SpaGCN, STAGATE, GraphST, BANKSY, Leiden, Louvain |
| Deconvolution | FlashDeconv, Cell2location, RCTD, DestVI, Stereoscope, SPOTlight, Tangram, CARD |
| Cell Communication | LIANA+, CellPhoneDB, CellChat (cellchat_r), FastCCC |
| Cell Type Annotation | Tangram, scANVI, CellAssign, mLLMCelltype, scType, SingleR |
| Differential Expression | Wilcoxon, t-test, Logistic Regression, pyDESeq2 |
| Trajectory & Velocity | CellRank, Palantir, DPT, scVelo, VeloVI |
| Spatial Statistics | Moran's I, Local Moran, Geary's C, Getis-Ord Gi*, Ripley's K, Co-occurrence, Neighborhood Enrichment, Centrality Scores, Local Join Count, Network Properties |
| Enrichment | GSEA, ORA, Enrichr, ssGSEA, Spatial EnrichMap |
| Spatial Genes | SpatialDE, SPARK-X, FlashS |
| Integration | Harmony, BBKNN, Scanorama, scVI |
| Other | CNV Analysis (InferCNVPy, Numbat), Spatial Registration (PASTE, STalign) |
Documentation
| Guide | Owns |
|---|---|
| Installation | Environment setup, package install, platform notes |
| Quick Start | First successful analysis after setup |
| Concepts | Method selection and analysis reasoning |
| Examples | Prompt recipes and workflow examples |
| Configuration | Exact MCP client configuration syntax |
| Troubleshooting | Symptom → fix guidance |
| Methods Reference | Canonical tool parameters and defaults |
| Full Docs | Complete documentation site |
Citation
If you use ChatSpatial in your research, please cite:
@article{Yang2026.02.26.708361,
author = {Yang, Chen and Zhang, Xianyang and Chen, Jun},
title = {ChatSpatial: Schema-Enforced Agentic Orchestration for Reproducible and Cross-Platform Spatial Transcriptomics},
elocation-id = {2026.02.26.708361},
year = {2026},
doi = {10.64898/2026.02.26.708361},
publisher = {Cold Spring Harbor Laboratory},
URL = {https://www.biorxiv.org/content/early/2026/03/01/2026.02.26.708361},
journal = {bioRxiv}
}
ChatSpatial orchestrates many excellent third-party methods. Please also cite the original tools your analysis used.
Contributing
Documentation improvements, bug reports, and new analysis methods are all welcome. See CONTRIBUTING.md.
Yorumlar (0)
Yorum birakmak icin giris yap.
Yorum birakSonuc bulunamadi







