years
Health Warn
- License — License: MIT
- Description — Repository has a description
- Active repo — Last push 0 days ago
- Low visibility — Only 5 GitHub stars
Code Warn
- Code scan incomplete — No supported source files were scanned during light audit
Permissions Pass
- Permissions — No dangerous permissions requested
No AI report is available for this listing yet.
Add years to your life. Your health record, as code. A personal longevity system built on Claude Code.
Years
Add years to your life. Your health record, as code.
Years is a personal longevity system built on Claude Code. Your DNA, bloodwork, scans, and visit notes live as markdown in a private git repo, with slash commands to organize and analyze them. You, your AI, and your doctor work from the same record.
Pre-alpha. Pair with a physician. Full caveats in §Status.
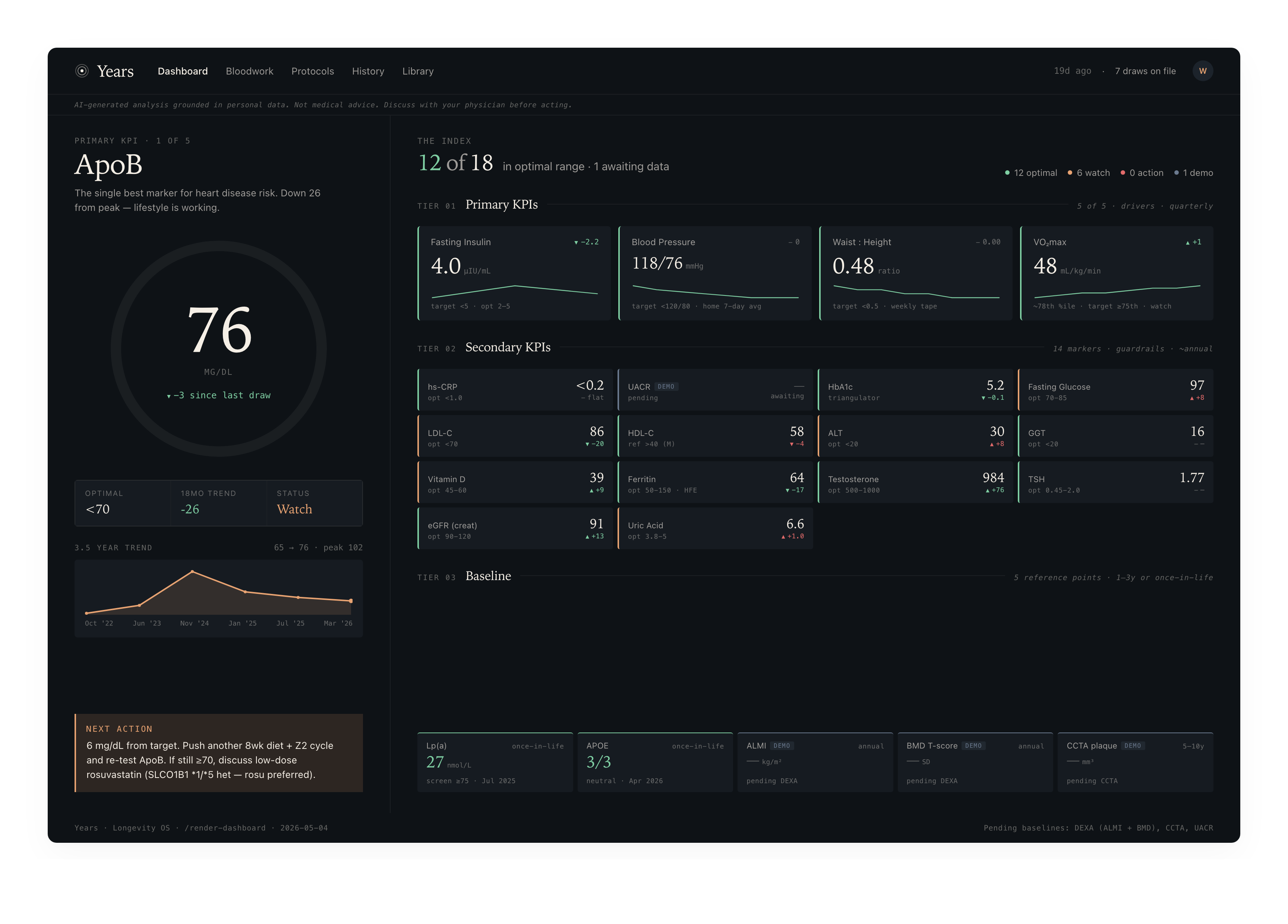
At a glance
- Open-source. No vendor lock-in and no hosted account that can vanish with your records.
- Markdown-as-database. Bloodwork, DNA, history, symptoms, and visits live as plain text on your machine. Readable by you, your physician, and your AI.
- Git-native. Your record is a private repo: diffable, branchable, backupable.
- Agent-native. No Node, Python, or build step. Slash commands are Markdown prompts Claude reads when you invoke them.
- Eight Claude Code slash commands:
/onboard— Set up your stack and capture starting context (identity, family history, next actions)./parse-bloodwork— Turn one lab PDF into a structured, tracked draw record./analyze-bloodwork— Read your latest draw through a longevity lens./catch-up-bloodwork— Process years of past lab PDFs into a single catch-up analysis./analyze-dna— Parse a 23andMe export against curated, actionable variants./check-in— Synthesize the next 3 highest-leverage moves across labs, screenings, and lifestyle./prep-appointment— Walk into your next clinician visit with the right questions./render-dashboard— Render your Primary KPIs as a self-contained HTML dashboard.
- Forkable evidence library. ~144 markers, 200+ DNA variants, ~15 domain syntheses.
Origin
Before my son was born, I asked my PCP what I should do to live well into my nineties. He told me I was healthy and to "come back in a year".
That answer bothered me. More than it probably should have.
So I started interrogating my own data: my biometrics, DNA, advanced bloodwork, and family history. Two things surfaced that routine care had missed.
The first was hemochromatosis, a genetic iron-overload condition. 23andMe flagged the potential. Iron labs came back elevated. A liver MRI confirmed iron overload. I've been able to manage through regular blood donation and reduced iron exposure.
The second was non-calcified soft plaque, caught on a CCTA. I'd ordered the scan after advanced lipid testing flagged risk that standard cholesterol panels had missed. A calcium score alone would have read zero and falsely reassured me. Now managed with a statin and lifestyle changes.
On most health measures I was in the top percentile and still trending toward premature heart disease and organ damage.
Years is the system I'm building to automate that process, and to help the next person figure out how to live a longer, healthier life.
What it's caught
Years answers "should I do X?", given your DNA, labs, imaging, and family history. Not "is X good in general?"
Three real questions Years has answered for me, all from the same record:
- Should I consider TRT? Almost certainly not as iron overload makes it a bad call.
- What should I add to my annual screening? Glaucoma screening, due to a
LOXL1polymorphism. - Which statin should I take? Rosuvastatin over atorvastatin, given myopathy risk from a
SLCO1B1variant.
None of these answers existed in any single chart, lab, or genetic report. They emerged from digging into DNA, bloodwork, imaging, family history, and daily biometrics together.
What you'll track
Years isn't a tracker. It's a tool for picking your next move: given where you actually are, what's the highest-leverage thing to do?
Some moves are cheap and high-leverage. A FIT test in your forties costs $50 and five minutes a year, and meaningfully cuts your colorectal cancer risk. Lifting your VO₂max takes 4–6 hours of zone-2 a week for months. Home BP monitoring is free, but mostly noise if you're already 115/75. What to do next depends on where you actually are.
Roughly, Years prioritizes by:
EV/cost ≈ (Δrisk × P(move it) × durability) / (hours + dollars + side effects + tail risk)
- Δrisk — how much an intervention shifts your long-term risk of dying or getting sick. Statins cut cardiovascular deaths by about 25%, regular exercise cuts overall death rates 20–30%, colorectal cancer screening cuts colorectal cancer deaths by roughly half.
- P(move it) — your honest probability of sticking with it. A protocol you won't sustain has near-zero expected value.
- Durability — separates one-time wins like hemochromatosis screening from things that decay the moment you stop, like sleep hygiene.
- Tail risk — the chance of a serious unknown harm. A statin has 30+ years of post-marketing data; a research peptide does not.
Primary KPIs are the five markers you're actively trying to move this cycle. Your five should reflect phenotype, family history, and what you've already addressed. A 60-year-old with confirmed plaque shouldn't be tracking the same five as a 30-year-old with a clean scan.
Years ships a default for adults with no special history: ApoB, fasting insulin / HOMA-IR, home blood pressure, waist-to-height ratio, and VO₂max trend from a smartwatch. Start here if you're starting from zero.
Otherwise, build your own five:
- Promote markers you need to move. Ferritin and TSAT if you're a hemochromatosis carrier, ApoB if your dad had a heart attack at 55, cystatin C eGFR if you're carrying a lot of muscle and standard creatinine eGFR is misleading.
- Demote markers reliably in your optimal range to the Secondary tier as guardrails, checked annually.
- Pick from the remaining candidates using the EV/cost lens above.
reference/panels.md §Phenotype-dependent guidance has more.
Secondary KPIs (guardrails and confirmatory readouts (~annual)). Markers that sit reliably in your optimal range serve as guardrails. Confirmatory readouts triangulate the Primary KPIs: HbA1c lags fasting insulin by months, hs-CRP catches inflammation that fasting insulin misses, UACR confirms blood pressure is actually reaching the kidney.
Baseline measurements (infrequent or once-in-life). DEXA every few years for bone density and body comp. CCTA for plaque burden when there's a reason to look. APOE and Lp(a) genotyping once, since neither changes.
How it's different
I've extracted my data from Forward, UCLA, Cedars, and a concierge physician. It was painful every time. A liver MRI of mine still lives on a CD in a drawer somewhere. I still can't tell you whether I got the Hep B vaccine.
Years is what I'm building so I don't have to do that again.
The record is a private git repo on your machine: plain Markdown files for your labs, DNA, imaging, and family history. Markers get flagged against optimal ranges, not lab-normal ones. And how a value moves across draws matters more than where it sits today. Most of the story is in the trend.
Claude Code reads the record through slash commands. Cross DNA against bloodwork. Bloodwork against family history. Ask the same question five different ways.
The reasoning rules live in reference/ as Markdown you fork. When new research breaks, anyone can update them. You can disagree with my defaults and override them in your own copy.
Quickstart
Prerequisites: Claude Code.
Clone years/ wherever you keep your projects. /onboard will create years-data/ next to it.
git clone https://github.com/[you]/years.git
cd years
claude
Then, inside Claude Code:
- Run
/onboard. It creates../years-data/, scaffolds the template, runs an intake (identity + family history + bloodwork/DNA status), and writes top-3 next-actions toyears-data/actions.md. Optionallygit initsyears-data/if you want it version-controlled. - Drop lab PDFs in
years-data/data/bloodwork/pdfs/(gitignored withinyears-data/— originals never leave your machine). - Single PDF: run
/parse-bloodwork <pdf>, then verify and edit the generated Markdown as needed. Multiple historical PDFs: run/catch-up-bloodworkto parse all and produce a recency-weighted analysis. - Run
/analyze-bloodwork(skip if you ran/catch-up-bloodwork— it produces an analysis inline).
years-data/ holds your personal records. Store it as a private git repo, plain folder, or an encrypted volume.
Don't have bloodwork or DNA results yet? Two baselines help: (1) a comprehensive bloodwork panel (Function Health, Superpower, or equivalent, see
reference/panels.md §Tier 1), (2) consumer DNA (23andMe etc., seereference/genetic-testing.md). Doing each at least once is generally helpful.
How it's organized
Years is two folders: this repo — the shared template (years/) — and your personal records (years-data/), which you keep private.
<wherever you cloned>/
├── years/ ← template, schemas, slash commands (public)
└── years-data/ ← your data (private)
Updates are one command: cd years && git pull. Your data is untouched; the pull operates only inside years/.
On every command, the agent reads three files first: the template CLAUDE.md (rules), your personal CLAUDE.md if it exists (your overrides), and your health_summary.md.
Inside years/ (the template — what you fork):
Schemas + slash commands. Directory structure, file conventions, the eight slash commands (full list in §Commands).
CLAUDE.md— reasoning rules, defaults, priority biomarkers, pharmacogenomic hard rules.reference/— the curated knowledge layer:ranges.md— ~144 markers with tighter flagging than lab "normal."dna-variants.md— 200+ curated rsIDs (risk alleles, confidence tiers, strand notes).panels.md— tiered testing strategy.<domain>.md— ~15 synthesis files spanning cardiovascular, iron, metabolic, hepatic, renal, hormonal, thyroid, inflammation, nutrients, sleep, training, nutrition, cancer, neurodegeneration.
Sourced from ~33 primary thinkers; contributions are citation-backed and reviewable.
Inside years-data/ (your records — what you create):
CLAUDE.md(yours, optional). Anything you add — your DNA results, the targets you're aiming for, notes about your body and history — overrides the defaults.health_summary.md— what's currently true about you: key theses, document map, imaging snapshot, family history.- Per-draw records, biomarker history, generated analyses, raw lab PDFs. Slash commands read inputs from here and write outputs back here, grounded in your variants, your trends, and cited evidence from
reference/.
Treat everything in years-data/ as personal health information. The included .gitignore only excludes raw PDFs (data/**/pdfs/).
One cloud touchpoint: Claude Code ships every /analyze-bloodwork context to Anthropic. Full boundary logic in CLAUDE.md §Invariants #2.
Status & trust
Pre-alpha. I've run a personal version of this for a few years, but all the commands, reference synthesis are new and in flux. It hasn't been reviewed by physicians and I'm not a professional in the health industry. Expect the thresholds, variants, and commands to change; wrong numbers, bad flags, and silent errors are possible. Expect breaking changes.
Much of the synthesis and prose was written with Claude and ChatGPT: reference files, variant rules, domain prose. Verify citations at the source. I'm not a physician; this is not medical advice. Read the analyses, check them against your raw PDFs, and work with a clinician. Don't act on anything you haven't verified.
What's next
- Wearable integration. Oura, Whoop, etc.
- Expanded DNA analysis. ClinVar quarterly intersection, PharmGKB drug-gene tables, polygenic scores, full-exome / WGS support beyond the curated rsID list.
- Imaging integration. CCTA, DEXA, etc.
- Symptom + lifestyle logging. Lightweight commands for nutrition, training, mood.
Commands
Slash commands are Markdown prompts that Claude Code picks up automatically; everything runs through Claude.
Typical day-1 path: /onboard → drop lab PDFs into ../years-data/data/bloodwork/pdfs/ → /catch-up-bloodwork → /analyze-dna <raw-file> (if you have one) → /check-in.
| Command | Argument | What it does |
|---|---|---|
/onboard |
— | Bootstraps a fresh stack. Creates ../years-data/, runs interactive intake, writes top-3 next-actions to actions.md. |
/parse-bloodwork |
<pdf> (optional; picker if omitted) |
Ingest a lab PDF into canonical Markdown. Only command that touches a PDF. |
/analyze-bloodwork |
latest | YYYY-MM-DD | filename-stem |
Two-pass synthesis — objective (shareable) + interpretive. Pass B may propose date-stamped appends to health_summary.md (per-edit confirmation, append-only). |
/catch-up-bloodwork |
— | One-shot catch-up for fresh forkers with multiple historical PDFs. Recency-weighted analysis. |
/analyze-dna |
<raw-file> |
Parse a 23andMe export against curated variants; bucket findings as Act-On-Now / Useful Context / Not Flagged / Not Assessable. Offers to append Act-On-Now findings to CLAUDE.md §Static genetics. |
/check-in |
— | Horizontal synthesis. Reads latest analysis + screenings + meds + lifestyle, writes a report to analysis/ + top-3 to actions.md. |
/render-dashboard |
— | Render the Longevity dashboard — 10 markers across three tiers (Primary KPIs + Secondary KPIs + Baseline measurements) — as a single self-contained HTML file (vanilla HTML/CSS/inline SVG, opens offline). |
/prep-appointment |
<visit-context> (free-text) |
Conversation scaffold for an upcoming clinician visit. |
/log-visit |
<slug | date | pdf-path> (optional) |
Post-visit capture. Converts a /prep-appointment future-visit file (or starts fresh), parses optional summary PDFs, proposes per-edit appends to conditions.md, medications-and-supplements.md, health_summary.md, and actions.md. |
Disclaimer
Years is a personal tooling template, not a medical product. Nothing here (reference/, generated analyses, or command output) is medical advice, diagnosis, or treatment. Use it as a thinking aid with a licensed clinician. Discuss any medication, supplement, dosing, or testing with your physician. No warranty of accuracy or fitness; no liability for decisions made from this content. In an emergency, call your local emergency number (911 in the US).
License
All code and infrastructure files (slash commands, schemas, templates, scripts) are MIT-licensed — see LICENSE.
Prose documentation in reference/ and other Markdown content is licensed under Creative Commons Attribution 4.0 — see LICENSE-CONTENT.
Contributions are accepted under the same terms (inbound = outbound; GitHub ToS §D.6). No CLA required.
Reviews (0)
Sign in to leave a review.
Leave a reviewNo results found