biocli
Health Gecti
- License — License: MIT
- Description — Repository has a description
- Active repo — Last push 0 days ago
- Community trust — 10 GitHub stars
Code Gecti
- Code scan — Scanned 12 files during light audit, no dangerous patterns found
Permissions Gecti
- Permissions — No dangerous permissions requested
This tool is an agent-first command-line interface designed for bioinformatics research. It allows users and AI agents to query multiple biological databases (like NCBI, UniProt, and KEGG), download datasets, and automatically prepare analysis-ready project directories.
Security Assessment
Overall Risk: Low
The rule-based code scan found no dangerous patterns across its files. It makes external network requests, but this is entirely expected given its core function of fetching data from public scientific databases. It does not request dangerous system permissions. No hardcoded secrets were detected, and accessing sensitive personal data is not a feature of this tool.
Quality Assessment
The project appears to be actively maintained, with its most recent code push occurring today. It uses the standard and highly permissive MIT license, which is excellent for open-source collaboration. Community trust is currently minimal but positive, represented by 10 GitHub stars. The documentation is thorough, clearly outlining its features, installation process, and how it compares to similar tools in the bioinformatics ecosystem.
Verdict
Safe to use.
Agent-first bioinformatics CLI: query NCBI/UniProt/KEGG/STRING/Ensembl/Enrichr, download data, prepare research-ready directories
biocli
Query biological databases from the terminal. Agent-first design.
biocli v0.3.8
NCBI · UniProt · KEGG · STRING · Ensembl · Enrichr
44 commands · 6 database backends · 10 workflow commands · 4 download commands
Install
npm install -g @yangfei_93sky/biocli
Requires Node.js >= 20. No API keys needed (optional NCBI key increases rate limit).
Why biocli
biocli is the only CLI that takes you from a research question to an analysis-ready working directory — scout datasets, download data, fetch annotations, all in one pipeline.
# Scout relevant datasets for your research question
biocli aggregate workflow-scout "TP53 breast cancer RNA-seq" --gene TP53
# Prepare a working directory with data + annotations + manifest
biocli aggregate workflow-prepare GSE315149 --gene TP53 --outdir ./project
Designed for AI agents (Claude Code, Codex CLI, etc.) — structured JSON output, per-command schema, self-describing help, batch input, local cache.
How biocli compares
| biocli | gget | BioMCP | EDirect | |
|---|---|---|---|---|
| Query biological databases | ✅ | ✅ | ✅ | ✅ |
| Structured JSON output | ✅ | ✅ | ✅ | ❌ |
| Cross-database aggregation | ✅ | ❌ | ✅ | ❌ |
| Download GEO/SRA data files | ✅ | ❌ | ❌ | ❌ |
| Dataset discovery (scout) | ✅ | ❌ | ❌ | ❌ |
| Working directory prep (prepare) | ✅ | ❌ | ❌ | ❌ |
| Agent command self-description | ✅ | ❌ | ⚠️ | ❌ |
| Safe preview (--plan/--dry-run) | ✅ | ❌ | ❌ | ❌ |
| Per-command JSON Schema | ✅ | ❌ | ❌ | ❌ |
| Local response cache | ✅ | ❌ | ❌ | ❌ |
| Batch input (--input) | ✅ | ❌ | ✅ | ✅ |
gget excels at sequence analysis (BLAST, AlphaFold, MUSCLE). BioMCP covers more biomedical entities (drugs, trials, diseases). EDirect has the deepest NCBI Entrez integration. biocli is the only one that combines query + download + data preparation into agent-orchestrated workflows.
Benchmark: Agent-First Biological Workflow Tasks (2026-04-04)
12 tasks across gene intelligence, variant interpretation, literature search, and data preparation. Task scores are automated from raw output; cross-cutting scores are manual audit with published justifications. Full methodology →
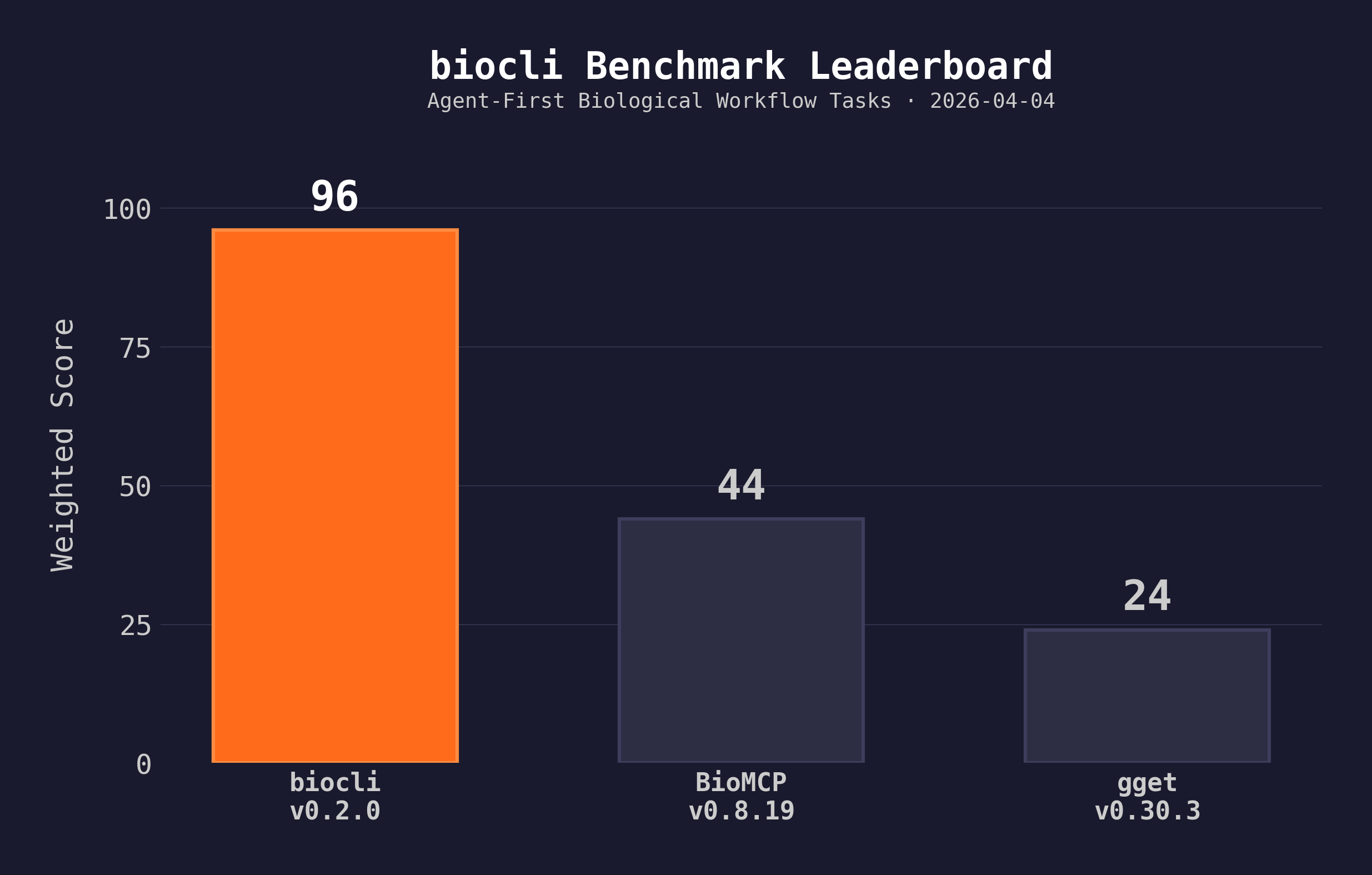
| Tool | Version | Task Success | Agent Readiness | Workflow Depth | Safety | Reproducibility | Total |
|---|---|---|---|---|---|---|---|
| biocli | 0.2.0 | 47/49 | 10/10 | 10/10 | 9/10 | 10/10 | 96/100 |
| BioMCP | 0.8.19 | 20/49 | 6/10 | 4/10 | 3/10 | 2/10 | 44/100 |
| gget | 0.30.3 | 8/49 | 3/10 | 2/10 | 2/10 | 1/10 | 24/100 |
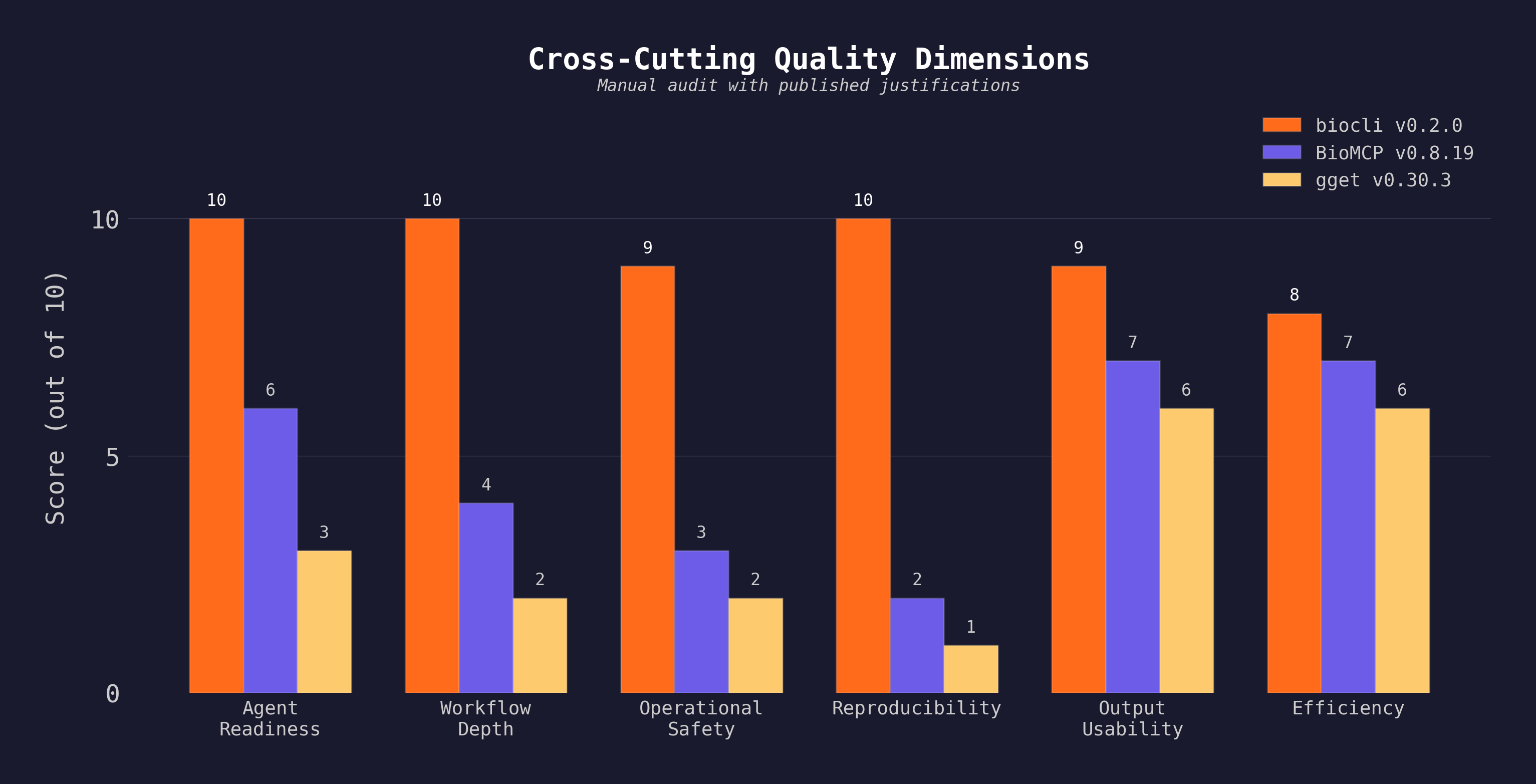
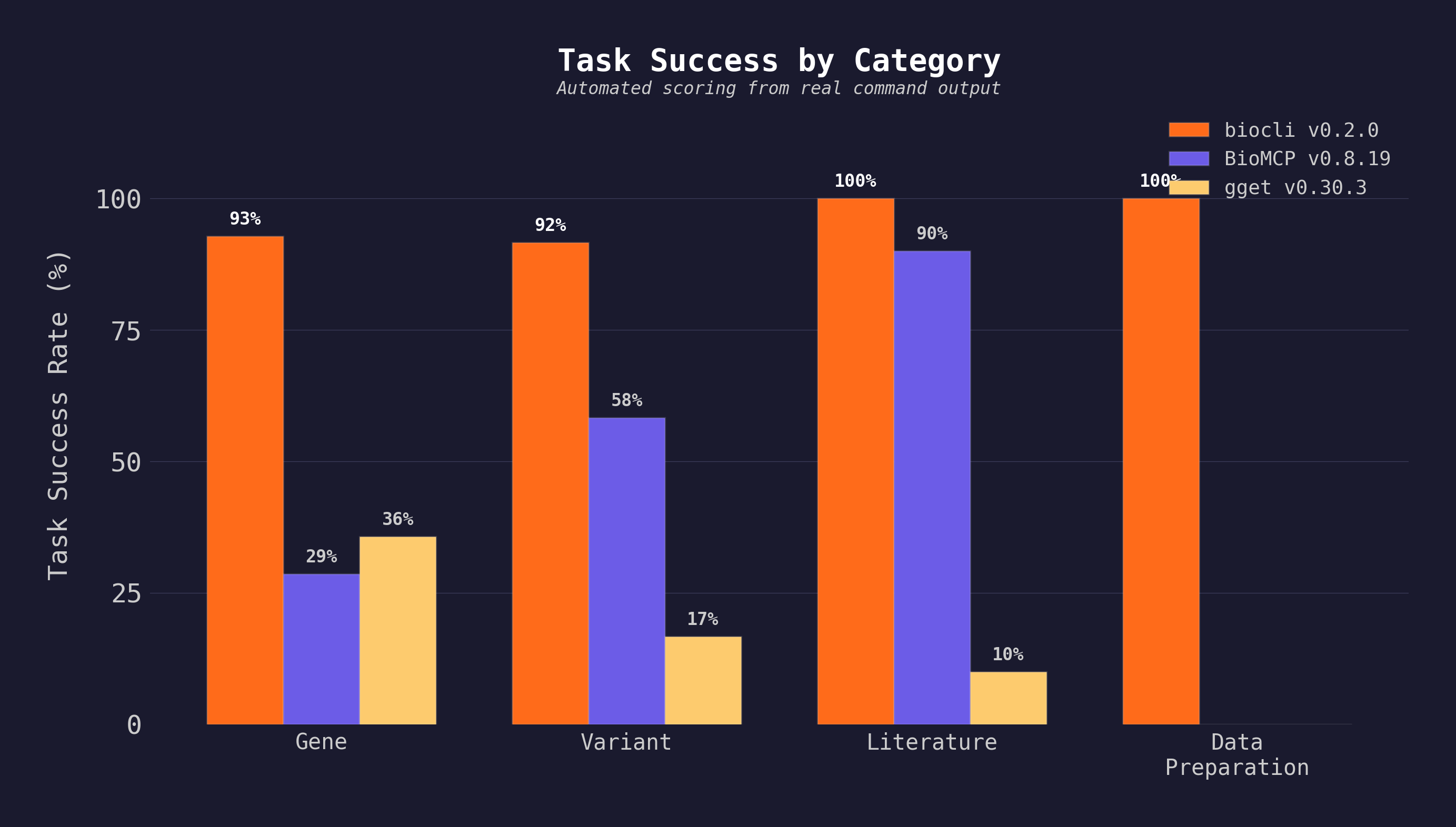
All three tools were installed (
npm install -g @yangfei_93sky/biocli,pip install gget==0.30.3,uv tool install biomcp-cli==0.8.19) and executed on the same machine with the same inputs. Raw stdout/stderr, scoring scripts, and runner scripts are inbenchmarks/. BioMCP excels at biomedical entity breadth (drugs, trials, diseases) not covered by this task set; gget excels at sequence analysis (BLAST, AlphaFold) not covered here.
Quick start
One command replaces 4 browser tabs:
biocli aggregate gene-dossier TP53 -f json
Returns a unified JSON with gene summary, protein function, KEGG pathways, GO terms, protein interactions, recent literature, and clinical variants — sourced from NCBI, UniProt, KEGG, STRING, PubMed, and ClinVar in parallel.
# Gene intelligence (NCBI + UniProt + KEGG + STRING + PubMed + ClinVar)
biocli aggregate gene-dossier TP53
# Variant interpretation (dbSNP + ClinVar + Ensembl VEP)
biocli aggregate variant-dossier rs334
# Literature review with abstracts
biocli aggregate literature-brief "CRISPR cancer immunotherapy" --limit 10
# Pathway enrichment (Enrichr + STRING)
biocli aggregate enrichment TP53,BRCA1,EGFR,MYC,CDK2
# Gene profile (NCBI + UniProt + KEGG + STRING)
biocli aggregate gene-profile TP53
All commands
Workflow commands (agent-optimized)
| Command | Sources | Use case |
|---|---|---|
aggregate gene-dossier <gene> |
NCBI+UniProt+KEGG+STRING+PubMed+ClinVar | Complete gene intelligence report |
aggregate variant-dossier <variant> |
dbSNP+ClinVar+Ensembl VEP | Variant interpretation |
aggregate variant-interpret <variant> |
dbSNP+ClinVar+VEP+UniProt | Variant interpretation with clinical context |
aggregate literature-brief <query> |
PubMed | Literature summary with abstracts |
aggregate enrichment <genes> |
Enrichr+STRING | Pathway/GO enrichment analysis |
aggregate gene-profile <gene> |
NCBI+UniProt+KEGG+STRING | Gene profile (no literature) |
aggregate workflow-scout <query> |
GEO+SRA | Scout datasets for a research question |
aggregate workflow-prepare <dataset> |
GEO+NCBI+UniProt+KEGG | Prepare research-ready directory with data + annotations |
aggregate workflow-annotate <genes> |
NCBI+UniProt+KEGG+Enrichr | Annotate gene list → genes.csv + pathways.csv + enrichment.csv + report.md |
aggregate workflow-profile <genes> |
NCBI+UniProt+KEGG+STRING+Enrichr | Gene set functional profile → shared pathways, interactions, GO terms |
Database commands (atomic)
| Database | Commands |
|---|---|
| PubMed | pubmed search, fetch, abstract, cited-by, related, info |
| Gene | gene search, info, fetch (FASTA download) |
| GEO | geo search, dataset, samples, download |
| SRA | sra search, run, download (FASTQ via ENA/sra-tools) |
| ClinVar | clinvar search, variant |
| SNP | snp lookup |
| Taxonomy | taxonomy lookup |
| UniProt | uniprot search, fetch, sequence (FASTA download) |
| KEGG | kegg pathway, link, disease, convert |
| STRING | string partners, network, enrichment |
| Ensembl | ensembl lookup, vep, xrefs |
| Enrichr | enrichr analyze |
Output formats
biocli gene info 7157 -f json # JSON (default for workflow commands)
biocli gene info 7157 -f table # Table (default for atomic commands)
biocli gene info 7157 -f yaml # YAML
biocli gene info 7157 -f csv # CSV
biocli gene info 7157 -f plain # Plain text
Agent-first result schema
All workflow commands (aggregate *) return a standard BiocliResult envelope:
{
"data": { ... },
"ids": { "ncbiGeneId": "7157", "uniprotAccession": "P04637", ... },
"sources": ["NCBI Gene", "UniProt", "KEGG", "STRING"],
"warnings": [],
"queriedAt": "2026-04-03T10:00:00.000Z",
"organism": "Homo sapiens",
"query": "TP53"
}
data— the actual result payloadids— cross-database identifiers for the queried entitysources— which databases contributed datawarnings— partial failures, ambiguous matches (never silently hidden)queriedAt— ISO timestamp for reproducibilityorganism— species context
Configuration
biocli config set api_key YOUR_NCBI_KEY # Optional: increases NCBI rate limit 3→10 req/s
biocli config set email [email protected]
biocli config show
Config stored at ~/.biocli/config.yaml.
Rate limits
| Database | Rate | Auth |
|---|---|---|
| NCBI | 3/s (10/s with API key) | Optional API key |
| UniProt | 50/s | None |
| KEGG | 10/s | None |
| STRING | 1/s | None |
| Ensembl | 15/s | None |
| Enrichr | 5/s | None |
All rate limits are enforced automatically per-database.
License
MIT
Yorumlar (0)
Yorum birakmak icin giris yap.
Yorum birakSonuc bulunamadi