SciNet
Health Warn
- License — License: MIT
- Description — Repository has a description
- Active repo — Last push 0 days ago
- Low visibility — Only 7 GitHub stars
Code Pass
- Code scan — Scanned 12 files during light audit, no dangerous patterns found
Permissions Pass
- Permissions — No dangerous permissions requested
This project provides an open-source Python client for interacting with the SciNet API. It is designed to help automate scientific research tasks, such as generating and evaluating academic ideas, predicting research trends, and profiling authors using a massive academic knowledge graph.
Security Assessment
Overall Risk: Low. The tool functions as an API client and makes external network requests to a hosted SciNet server to retrieve data. The automated code scan of 12 files found no dangerous patterns, no hardcoded secrets, and no risky shell command executions. Users should simply be aware that the inputs and generated research requests are sent externally to the third-party API.
Quality Assessment
The project is actively maintained, with its last push occurring today. It is released under the permissive and standard MIT license, making it highly accessible for developers. However, it currently has extremely low community visibility with only 7 GitHub stars, indicating it is likely a new or niche academic tool without a broad established user base to validate it.
Verdict
Safe to use, though developers should keep in mind that all task executions and queries are processed via an external hosted API.
A Large-Scale Knowledge Graph for Automated Scientific Research
SciNet: A Large-Scale Knowledge Graph for Automated Scientific Research
Open-source client for running literature-grounded scientific research tasks on top of SciNet API.
📑 Table of Contents
✨ Overview
SciNet is a large-scale, multi-disciplinary, heterogeneous academic resource knowledge graph designed as a panoramic scientific evolution network. By integrating over 43M papers from 26 disciplines, and a total of 157M entites and 3B triplets, SciNet provides a structured topological cognitive substrate that dismantles disciplinary barriers and furnishes AI agents with a global perspective.
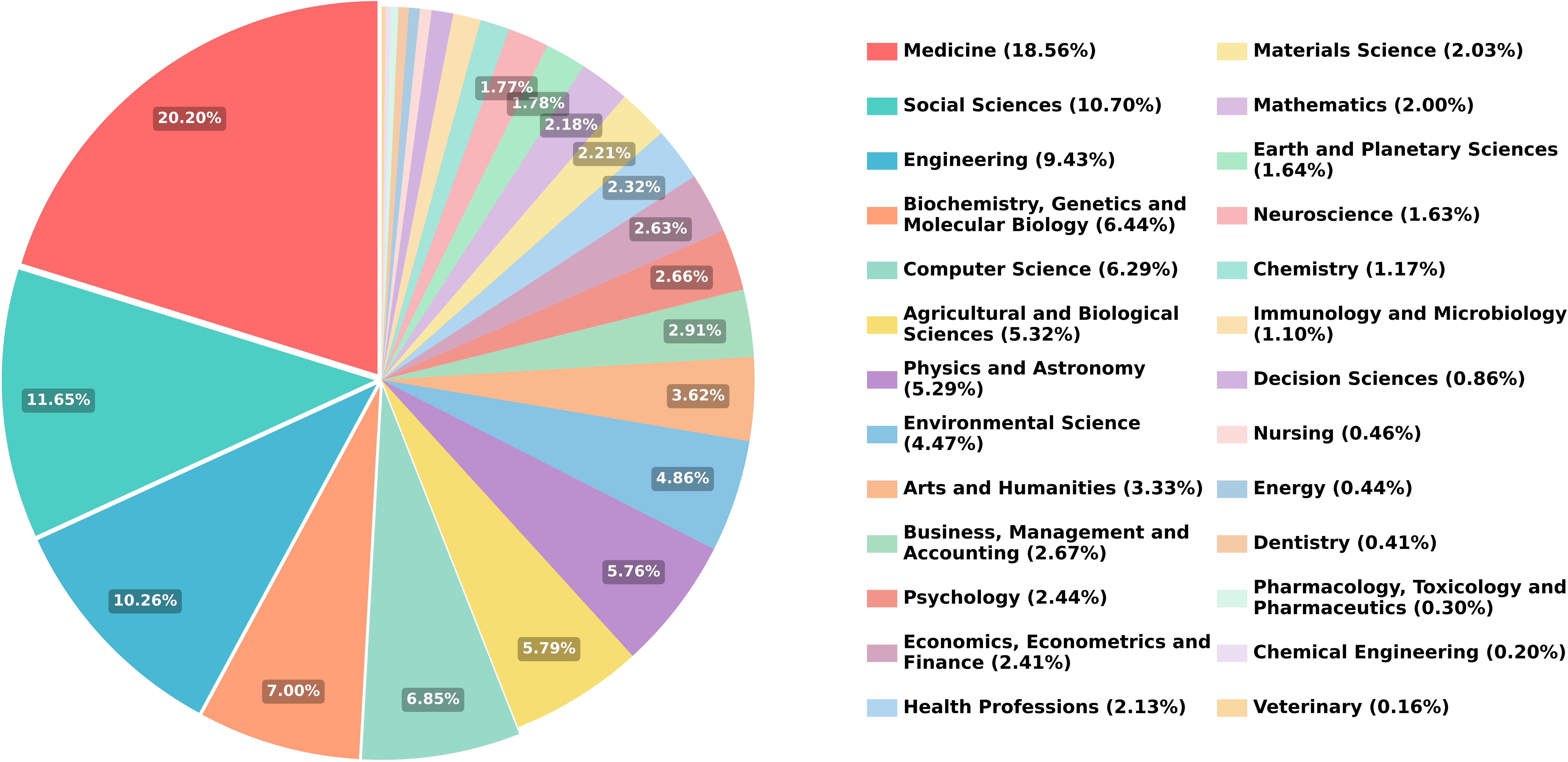
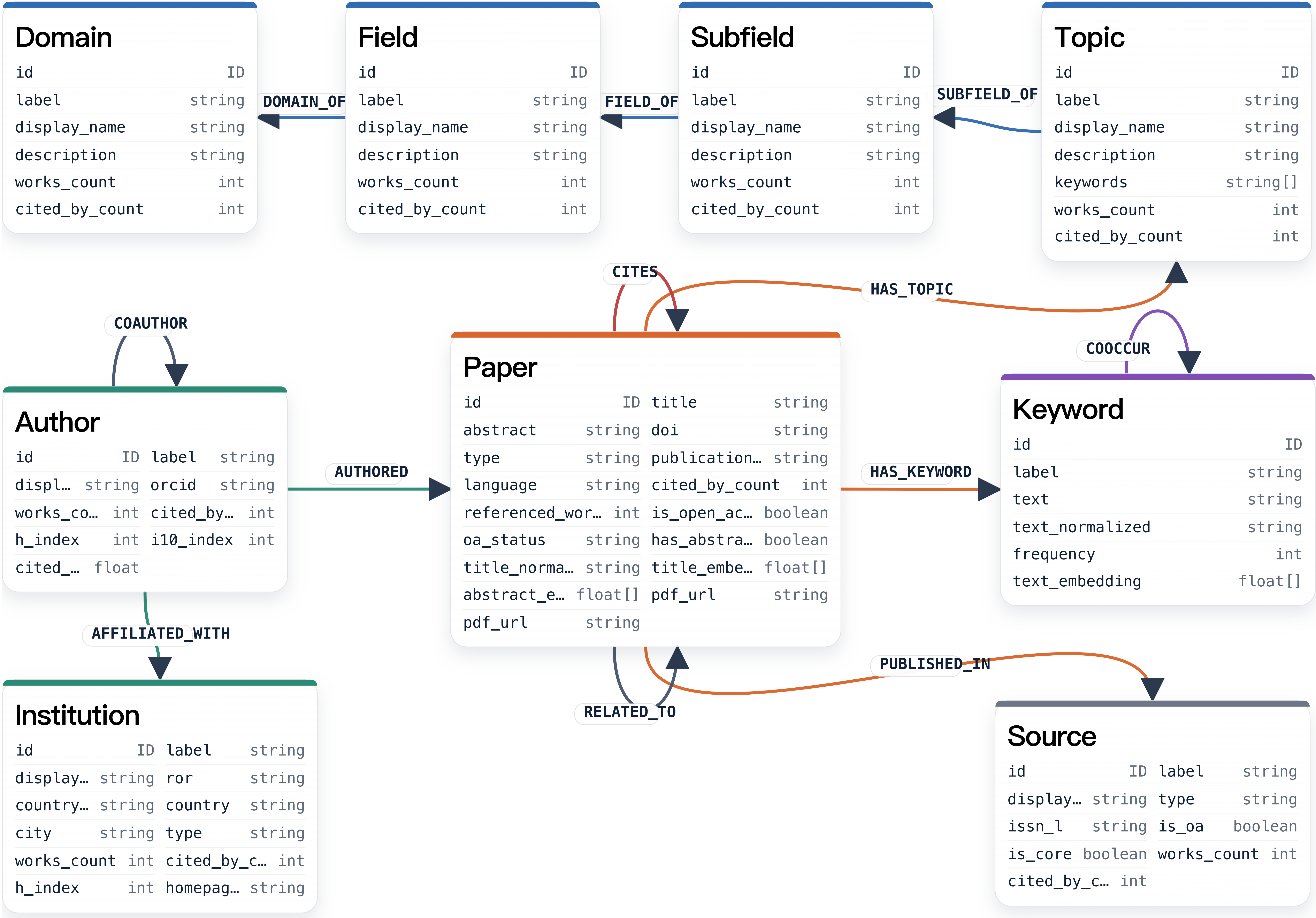
This repository provides a runnable client for several scientific research workflows, including idea evaluation, topic review, author discovery, author profiling, and idea generation.
Each run is driven by CLI inputs plus optional runtime parameter overrides. The client also writes a request.json file into the run directory so every execution remains easy to inspect and reproduce later.
The local client is responsible for:
- building a structured request
- calling a hosted
SciNet API - running client-side post-processing such as reranking, PDF parsing, grounding, and Markdown report generation
Users do not need to connect to Neo4j or other database components directly.
🧩 Supported Tasks
| Task Type | Required Input | Main Output |
|---|---|---|
Idea Grounding and Evaluation |
--idea-text or --pdf-path |
grounded evidence, paragraph matches, and idea-level analysis |
Idea Generation |
--topic-text |
generated ideas grounded in retrieved literature |
Research Trend Predicting |
--topic-text |
topic evolution summary and representative papers |
Related Author Retrieval |
--idea-text or --pdf-path |
related authors and supporting papers |
Researcher Background Review |
--author-name |
research trajectory and representative works |
🛠️ GROBID
GROBID extracts structured metadata from scientific PDFs, including titles, authors, abstracts, and references.
GROBID is needed for:
Idea Grounding and EvaluationRelated Author Retrievalwhen using--pdf-path
Example startup with Docker:
docker pull lfoppiano/grobid:latest
docker run -d --rm --name grobid -p 8070:8070 lfoppiano/grobid:latest
curl http://127.0.0.1:8070/api/isalive
📂 Layout
This repository is a lightweight client for SciNet workflows.
run_scinet.py: main entrypoint for runsscinet/: main runtime package, including CLI handling, task dispatch, retrieval, grounding, and Markdown renderingreferences/search/: reference and demonstration code for the search logic; it is kept for inspection and illustration, and is not part of the default runtime path
🚀 Quick Start
Use the following steps to get a working run from a clean checkout.
1. Create an environment and install dependencies
python3 -m venv .venv
source .venv/bin/activate
pip install -U pip
pip install -r requirements.txt
2. Create the environment file
cp .env.example .env
We currently provide a public SciNet API endpoint for community use. Fill in the required variables:
SCINET_API_BASE_URL=http://scinet.openkg.cn
SCINET_API_KEY=scinet-public-key
LLM_PROVIDER=openai_compatible
LLM_API_KEY=replace-me
LLM_BASE_URL=https://your-openai-compatible-endpoint/v1
LLM_MODEL=your-model-name
# Legacy compatibility keys. New setups should prefer LLM_*.
OPENAI_API_KEY=replace-me
OPENAI_BASE_URL=https://your-openai-compatible-endpoint/v1
OPENAI_MODEL=your-model-name
GROBID_BASE_URL=http://127.0.0.1:8070
OA_API_KEY=
OPENALEX_MAILTO=
You can get an OpenAlex API key from here.
Required variables:
| Variable | Required For | Notes |
|---|---|---|
SCINET_API_BASE_URL |
all tasks | hosted SciNet API base URL |
SCINET_API_KEY |
all tasks | sent as X-API-Key |
LLM_PROVIDER |
all tasks | provider selector, currently openai_compatible |
LLM_API_KEY |
all tasks | used for planning, reranking, and summarization |
LLM_BASE_URL |
all tasks | OpenAI-compatible base URL |
LLM_MODEL |
all tasks | chat model name |
GROBID_BASE_URL |
PDF tasks | needed for --pdf-path flows |
OA_API_KEY |
optional | OpenAlex fallback support |
OPENALEX_MAILTO |
optional | OpenAlex contact email |
3. Make sure the required services are ready
- a hosted
SciNet API - an LLM endpoint exposed in OpenAI-compatible format
- GROBID if you want to use
--pdf-path
4. Run a task
python3 run_scinet.py \
--task-type "Research Trend Predicting" \
--topic-text "research idea evaluation with large language models" \
--pretty
5. Check the output
Each run creates a directory under runs/ containing:
request.jsonresult.jsonresult.md
🧪 Run Tasks
Idea Grounding and Evaluation
python3 run_scinet.py \
--task-type "Idea Grounding and Evaluation" \
--idea-text "Use literature-grounded evidence to evaluate research ideas." \
--pretty
With PDF input:
python3 run_scinet.py \
--task-type "Idea Grounding and Evaluation" \
--pdf-path /absolute/path/to/paper.pdf \
--params-json '{"search_final_top_k": 15, "manifest_top_k": 10}' \
--pretty
Idea Generation
python3 run_scinet.py \
--task-type "Idea Generation" \
--topic-text "scientific idea generation with retrieval-augmented large language models" \
--pretty
Research Trend Predicting
python3 run_scinet.py \
--task-type "Research Trend Predicting" \
--topic-text "research idea evaluation with large language models" \
--pretty
Related Author Retrieval
python3 run_scinet.py \
--task-type "Related Author Retrieval" \
--idea-text "knowledge-grounded evaluation of scientific research ideas" \
--pretty
Researcher Background Review
python3 run_scinet.py \
--task-type "Researcher Background Review" \
--author-name "Geoffrey Hinton" \
--pretty
You can also override task parameters from your own JSON file:
python3 run_scinet.py \
--task-type "Idea Grounding and Evaluation" \
--idea-text "Use literature-grounded evidence to evaluate research ideas." \
--params-file /absolute/path/to/params.json \
--pretty
For Idea Grounding and Evaluation, model-related overrides can be supplied through --params-file or --params-json.
You can also set local model paths once in .env:
SCINET_EMBEDDING_MODEL_PATH=/absolute/path/to/BAAI--bge-large-en-v1.5
SCINET_RERANKER_MODEL_PATH=/absolute/path/to/BAAI--bge-reranker-large
grounded_review also accepts query_provider, query_model, and query_api_url overrides in params.
If omitted, it resolves them from LLM_PROVIDER, LLM_MODEL, and LLM_BASE_URL.
By default, Idea Grounding and Evaluation uses:
- embedding model:
BAAI/bge-large-en-v1.5huggingface_url - reranker model:
BAAI/bge-reranker-largehuggingface_url
The first run may download these models into the local Hugging Face cache.
📝 TODO
- CLI Tools. Add more user-facing CLI capabilities so downstream users and AI agents can invoke retrieval workflows without touching database internals.
- Skills. Package reusable agent skills for common scientific discovery workflows and expose best practices as easier-to-load components.
- More Knowledge. Integrate more knowledge forms beyond paper-centric entities, such as datasets, code, standards, theorems, and experimental experience.
- Benchmark and Evaluation. Build dedicated benchmarks and evaluation protocols for downstream scientific research tasks supported by SciNet.
- Dynamic Update. Improve dynamic knowledge updates toward a more systematic and frequent refresh mechanism.
✍️ Citation
If you find our work helpful, please use the following citations.
License
This project is licensed under the MIT License - see the LICENSE file for details.
Reviews (0)
Sign in to leave a review.
Leave a reviewNo results found


